Identification of INHBA as a potential biomarker for gastric cancer through a comprehensive analysis
- PMID: 37528145
- PMCID: PMC10394090
- DOI: 10.1038/s41598-023-39784-1
Identification of INHBA as a potential biomarker for gastric cancer through a comprehensive analysis
Abstract
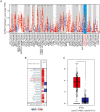
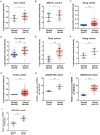
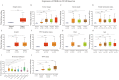
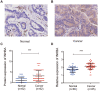
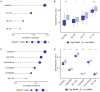
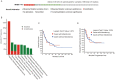
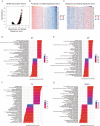
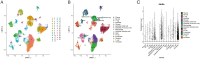
Inhibin subunit beta A (INHBA) is a member of the transforming growth factor-beta (TGF-β) superfamily that plays a fundamental role in various cancers. However, a systematic analysis of the exact role of INHBA in patients with gastric cancer (GC) has not yet been conducted. We evaluated the expression levels of INHBA and the correlation between INHBA and GC prognosis in GC. The relationship between INHBA expression, immune infiltration levels, and type markers of immune cells in GC was also explored. In addition, we studied INHBA mutations, promoter methylation, and functional enrichment analysis. Besides, high expression levels of INHBA in GC were significantly related to unfavorable prognosis. INHBA was negatively correlated with B cell infiltration, but positively correlated with macrophage and most anticancer immunity steps. INHBA expression was positively correlated with the type markers of CD8+ T cells, neutrophils, macrophages, and dendritic cells. INHBA has a weak significant methylation level change between tumor and normal tissues and mainly enriched in cancer-related signaling pathways. The present study implies that INHBA may serve as a potential biomarker for predicting the prognosis of patients with GC. INHBA is a promising predictor of immunotherapy response, with higher levels of INHBA indicating greater sensitivity.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures







Similar articles
-
Comprehensive analysis of inhibin-β A as a potential biomarker for gastrointestinal tract cancers through bioinformatics approaches.Sci Rep. 2025 Jan 7;15(1):1090. doi: 10.1038/s41598-024-72679-3. Sci Rep. 2025. PMID: 39774945 Free PMC article.
-
INHBA, transcriptionally activated by SPI1, facilitates gastric cancer progression by inducing macrophage recruitment and M2 polarization via activating the TGF-β signaling to increase CCL2.Pathol Res Pract. 2025 May;269:155920. doi: 10.1016/j.prp.2025.155920. Epub 2025 Mar 19. Pathol Res Pract. 2025. PMID: 40132395
-
DNA hypomethylation of INHBA promotes tumor progression and predicts prognosis and immune status of gastric cancer.Hereditas. 2024 Nov 14;161(1):45. doi: 10.1186/s41065-024-00347-7. Hereditas. 2024. PMID: 39543755 Free PMC article.
-
INHBA gene silencing inhibits gastric cancer cell migration and invasion by impeding activation of the TGF-β signaling pathway.J Cell Physiol. 2019 Aug;234(10):18065-18074. doi: 10.1002/jcp.28439. Epub 2019 Apr 8. J Cell Physiol. 2019. PMID: 30963572
-
Inflammation-related factors predicting prognosis of gastric cancer.World J Gastroenterol. 2014 Apr 28;20(16):4586-96. doi: 10.3748/wjg.v20.i16.4586. World J Gastroenterol. 2014. PMID: 24782611 Free PMC article. Review.
Cited by
-
Comprehensive analysis of inhibin-β A as a potential biomarker for gastrointestinal tract cancers through bioinformatics approaches.Sci Rep. 2025 Jan 7;15(1):1090. doi: 10.1038/s41598-024-72679-3. Sci Rep. 2025. PMID: 39774945 Free PMC article.
-
Identification and validation of serum MUC17 as a non-invasive early warning biomarker for screening of gastric intraepithelial neoplasia.Transl Oncol. 2025 Jan;51:102207. doi: 10.1016/j.tranon.2024.102207. Epub 2024 Nov 23. Transl Oncol. 2025. PMID: 39580962 Free PMC article.
-
Predictive Factors of Immunotherapy in Gastric Cancer: A 2024 Update.Diagnostics (Basel). 2024 Jun 13;14(12):1247. doi: 10.3390/diagnostics14121247. Diagnostics (Basel). 2024. PMID: 38928662 Free PMC article. Review.
-
Molecular insights into programmed cell death in esophageal squamous cell carcinoma.PeerJ. 2024 Jul 10;12:e17690. doi: 10.7717/peerj.17690. eCollection 2024. PeerJ. 2024. PMID: 39006030 Free PMC article.
-
Serum basic fibroblast growth factor and interleukin-1β predict the effect of first-line chemotherapy in patients with advanced gastric cancer.World J Clin Cases. 2023 Sep 16;11(26):6083-6090. doi: 10.12998/wjcc.v11.i26.6083. World J Clin Cases. 2023. PMID: 37731570 Free PMC article.
References
-
- Tan, P. & Yeoh, K. G. Genetics and molecular pathogenesis of gastric adenocarcinoma. Gastroenterology149, 1153–1162e1153. 10.1053/j.gastro.2015.05.059 (2015). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

