Genomics of Acinetobacter baumannii iron uptake
- PMID: 37549061
- PMCID: PMC10483418
- DOI: 10.1099/mgen.0.001080
Genomics of Acinetobacter baumannii iron uptake
Abstract
Iron is essential for growth in most bacteria due to its redox activity and its role in essential metabolic reactions; it is a cofactor for many bacterial enzymes. The bacterium
Keywords: Acinetobacter baumannii; haem; iron uptake; phylogeny; sequence type; siderophore.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures
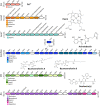


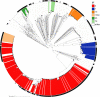
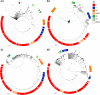
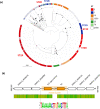

Similar articles
-
Acinetobacter baumannii can use multiple siderophores for iron acquisition, but only acinetobactin is required for virulence.PLoS Pathog. 2020 Oct 19;16(10):e1008995. doi: 10.1371/journal.ppat.1008995. eCollection 2020 Oct. PLoS Pathog. 2020. PMID: 33075115 Free PMC article.
-
Dissemination of novel Tn7 family transposons carrying genes for synthesis and uptake of fimsbactin siderophores among Acinetobacter baumannii isolates.Microb Genom. 2021 Mar;7(3):mgen000548. doi: 10.1099/mgen.0.000548. Epub 2021 Mar 22. Microb Genom. 2021. PMID: 33749577 Free PMC article.
-
Acinetobactin Isomerization Enables Adaptive Iron Acquisition in Acinetobacter baumannii through pH-Triggered Siderophore Swapping.ACS Infect Dis. 2016 Feb 12;2(2):157-68. doi: 10.1021/acsinfecdis.5b00145. Epub 2015 Dec 23. ACS Infect Dis. 2016. PMID: 27624967
-
Iron Acquisition Mechanisms and Their Role in the Virulence of Acinetobacter baumannii.Infect Immun. 2022 Oct 20;90(10):e0022322. doi: 10.1128/iai.00223-22. Epub 2022 Sep 6. Infect Immun. 2022. PMID: 36066263 Free PMC article. Review.
-
Current biochemical understanding regarding the metabolism of acinetobactin, the major siderophore of the human pathogen Acinetobacter baumannii, and outlook for discovery of novel anti-infectious agents based thereon.Nat Prod Rep. 2020 Apr 1;37(4):477-487. doi: 10.1039/c9np00046a. Epub 2019 Oct 29. Nat Prod Rep. 2020. PMID: 31661538 Review.
Cited by
-
Comparative genomics of Acinetobacter baumannii from Egyptian healthcare settings reveals high-risk clones and resistance gene mobilization.BMC Infect Dis. 2025 Jun 11;25(1):803. doi: 10.1186/s12879-025-11185-x. BMC Infect Dis. 2025. PMID: 40500700 Free PMC article.
-
Construction and Validation of a Predictive Model for Mortality Risk in Patients with Acinetobacter baumannii Bloodstream Infection.Infect Drug Resist. 2024 Nov 26;17:5247-5260. doi: 10.2147/IDR.S491537. eCollection 2024. Infect Drug Resist. 2024. PMID: 39624638 Free PMC article.
-
Expanding the substrate selectivity of the fimsbactin biosynthetic adenylation domain, FbsH.bioRxiv [Preprint]. 2024 Jul 26:2024.07.26.605314. doi: 10.1101/2024.07.26.605314. bioRxiv. 2024. Update in: ACS Chem Biol. 2024 Dec 20;19(12):2451-2461. doi: 10.1021/acschembio.4c00512. PMID: 39091846 Free PMC article. Updated. Preprint.
-
Positron Emission Tomography Imaging of Acinetobacter baumannii Infection: Comparison of Gallium-68 Labeled Siderophores.ACS Infect Dis. 2025 Apr 11;11(4):917-928. doi: 10.1021/acsinfecdis.4c00946. Epub 2025 Mar 18. ACS Infect Dis. 2025. PMID: 40099411 Free PMC article.
-
Exploring the clinical outcomes and molecular characteristics of Acinetobacter baumannii bloodstream infections: a study of sequence types, capsular types, and drug resistance in China.Front Cell Infect Microbiol. 2025 Feb 17;15:1549940. doi: 10.3389/fcimb.2025.1549940. eCollection 2025. Front Cell Infect Microbiol. 2025. PMID: 40034394 Free PMC article.
References
-
- CDC Atlanta, GA: U.S. Department of Health and Human Services; 2019. Antibiotic Resistance Threats in the United States, 2019. 10.15620/cdc:82532.
-
- Food and Drug Administration Establishing a list of qualifying pathogens under the food and Drug Administration safety and innovation act. Fed Register. 2013;78:35155–35173. - PubMed
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

