Eosinophil Phenotypes Are Functionally Regulated by Resolvin D2 during Allergic Lung Inflammation
- PMID: 37552821
- PMCID: PMC10704125
- DOI: 10.1165/rcmb.2023-0121OC
Eosinophil Phenotypes Are Functionally Regulated by Resolvin D2 during Allergic Lung Inflammation
Abstract
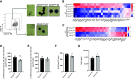
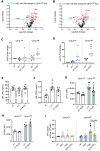
Eosinophils (Eos) reside in multiple organs during homeostasis and respond rapidly to an inflammatory challenge. Although Eos share chemical staining properties, they also demonstrate phenotypic and functional plasticity that is not fully understood. Here, we used a murine model of allergic lung inflammation to characterize Eos subsets and determine their spatiotemporal and functional regulation during inflammation and its resolution in response to resolvin D2 (RvD2), a potent specialized proresolving mediator. Two Eos subsets were identified by CD101 expression with distinct anatomic localization and transcriptional signatures at baseline and during inflammation. CD101low Eos were predominantly located in a lung vascular niche and responded to allergen challenge by moving into the lung interstitium. CD101high Eos were predominantly located in bronchoalveolar lavage (BAL) and extravascular lung, only present during inflammation, and had transcriptional evidence for cell activation. RvD2 reduced total Eos numbers and changed their phenotype and activation by at least two distinct mechanisms: decreasing interleukin 5-dependent recruitment of CD101low Eos and decreasing conversion of CD101low Eos to CD101high Eos. Collectively, these findings indicate that Eos are a heterogeneous pool of cells with distinct activation states and spatiotemporal regulation during resolution of inflammation and that RvD2 is a potent proresolving mediator for Eos recruitment and activation.
Keywords: allergic inflammation; eosinophils; resolvin D2; specialized proresolving mediators.
Figures



Comment in
-
Back to the Eosinophil: Resolvin Spatiotemporal Regulation.Am J Respir Cell Mol Biol. 2023 Dec;69(6):608-609. doi: 10.1165/rcmb.2023-0261ED. Am J Respir Cell Mol Biol. 2023. PMID: 37703390 Free PMC article. No abstract available.
References
-
- Wechsler ME, Munitz A, Ackerman SJ, Drake MG, Jackson DJ, Wardlaw AJ, et al. Eosinophils in health and disease: a state-of-the-art review. Mayo Clin Proc . 2021;96:2694–2707. - PubMed
-
- Fanta CH. Asthma. N Engl J Med . 2009;360:1002–1014. - PubMed
-
- Kay AB. Paul Ehrlich and the early history of granulocytes. Microbiol Spectr . 2016;4 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

