Comprehensive analysis of circRNAs for N7-methylguanosine methylation modification in human oral squamous cell carcinoma
- PMID: 37554544
- PMCID: PMC10405248
- DOI: 10.1096/fba.2023-00036
Comprehensive analysis of circRNAs for N7-methylguanosine methylation modification in human oral squamous cell carcinoma
Abstract
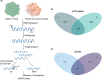
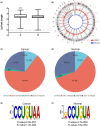
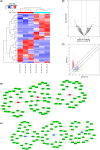
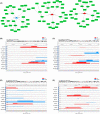
N7-methylguanosine (m7G) modification is closely related to the occurrence of tumors. However, the m7G modification of circRNAs in oral squamous cell carcinoma (OSCC) remains to be investigated. Methylated RNA immunoprecipitation sequencing (MeRIP-seq) was used to measure the methylation levels of m7G and identify m7G sites in circRNAs in human OSCC and normal tissues. The host genes of differentially methylated and differentially expressed circRNAs were analyzed by Gene Ontology (GO) enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses, and circRNA-miRNA-mRNA networks were predicted using the miRanda and miRDB databases. The analysis identified 2348 m7G peaks in 624 circRNAs in OSCC tissues. In addition, the source of m7G-methylated circRNAs in OSCC was mainly the sense overlap region compared with normal tissues. The most conserved m7G motif in OSCC tissues was CCUGU, whereas the most conserved motif in normal tissues was RCCUG (R = G/A). Importantly, GO enrichment and KEGG pathway analysis showed that the host genes of differentially methylated and differentially expressed circRNAs were involved in many cellular biological functions. Furthermore, the significantly differentially expressed circRNAs were analyzed to predict the circRNA-miRNA-mRNA networks. This study revealed the whole profile of circRNAs of differential m7G methylation in OSCC and suggests that m7G-modified circRNAs may impact the development of OSCC.
Keywords: N7‐methylguanosine (m7G); RNA methylation; circular RNA; methylated RNA immunoprecipitation sequencing (MeRIP‐seq); oral squamous cell carcinoma (OSCC).
© 2023 The Authors. FASEB BioAdvances published by Wiley Periodicals LLC on behalf of The Federation of American Societies for Experimental Biology.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



References
-
- Ali J, Sabiha B, Jan HU, Haider SA, Khan AA, Ali SS. Genetic etiology of oral cancer. Oral Oncol. 2017;70:23‐28. - PubMed
-
- Shahinas J, Hysi D. Methods and risk of bias in molecular marker prognosis studies in oral squamous cell carcinoma. Oral Dis. 2018;24:115‐119. - PubMed
-
- Sung H, Ferlay J, Siegel RL, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209‐249. - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
