Mettl3 synergistically regulates TGF-β/SMAD2/3 to promote proliferation and metastasis of gastric cancer
- PMID: 37560008
- PMCID: PMC10408465
Mettl3 synergistically regulates TGF-β/SMAD2/3 to promote proliferation and metastasis of gastric cancer
Abstract
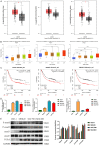
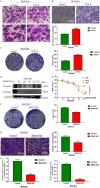
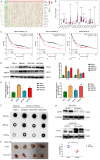
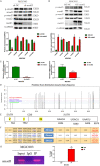
Transforming Growth factor-β (TGF-β)/Smad signaling is a complex regulatory network that both inhibits and promotes tumorigenesis. However, the mechanisms underlying the function of TGF-β/Smad signaling pathway remain to be fully elucidated. As a methyltransferase, METTL3 is closely related to tumor development, but the role of METTL3 in the proliferation and metastasis of TGF-β/Smad-activated gastric cancer (GC) is unclear. In this study, we identified TGF-β/Smad2/3 axis as an important carcinogenic pathway in GC, which significantly promoted the proliferation and metastasis of GC. Furthermore, we found that Smad3 mRNA could be modified by m6A, which was subsequently recognized and stabilized by IGF2BP2, thereby enhancing Smad3 protein expression and promoting the activation of TGF-β/Smad pathway. Importantly, we also found that METTL3 could combine with p-Smad3 to regulate the transcription of downstream target genes. Therefore, this study revealed a novel mechanism by which METTL3 synergistically regulates TGF-β/Smad2/3 signaling and provide a new potential therapeutic target for the treatment of GC.
Keywords: Gastric cancer; METTL3; TGF-β/Smads; interaction.
AJCR Copyright © 2023.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



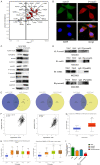
References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
