Structural basis of microtubule depolymerization by the kinesin-like activity of HIV-1 Rev
- PMID: 37572662
- PMCID: PMC10592302
- DOI: 10.1016/j.str.2023.07.009
Structural basis of microtubule depolymerization by the kinesin-like activity of HIV-1 Rev
Abstract
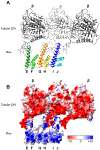
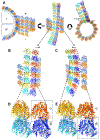
HIV-1 Rev is an essential regulatory protein that transports unspliced and partially spliced viral mRNAs from the nucleus to the cytoplasm for the expression of viral structural proteins. During its nucleocytoplasmic shuttling, Rev interacts with several host proteins to use the cellular machinery for the advantage of the virus. Here, we report the 3.5 Å cryo-EM structure of a 4.8 MDa Rev-tubulin ring complex. Our structure shows that Rev's arginine-rich motif (ARM) binds to both the acidic surfaces and the C-terminal tails of α/β-tubulin. The Rev-tubulin interaction is functionally homologous to that of kinesin-13, potently destabilizing microtubules at sub-stoichiometric levels. Expression of Rev in astrocytes and HeLa cells shows that it can modulate the microtubule cytoskeleton within the cellular environment. These results show a previously undefined regulatory role of Rev.
Keywords: CTT; HIV-1; Microtubules; Rev; Rings; Tubulin.
Published by Elsevier Ltd.
Conflict of interest statement
Declaration of interests The authors declare that they have no conflicts of interest with the contents of this article.
Figures


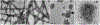
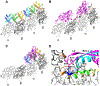



Similar articles
-
Nuclear Export Signal Masking Regulates HIV-1 Rev Trafficking and Viral RNA Nuclear Export.J Virol. 2017 Jan 18;91(3):e02107-16. doi: 10.1128/JVI.02107-16. Print 2017 Feb 1. J Virol. 2017. PMID: 27852860 Free PMC article.
-
HIV-1 rev depolymerizes microtubules to form stable bilayered rings.J Cell Biol. 2000 Jul 24;150(2):349-60. doi: 10.1083/jcb.150.2.349. J Cell Biol. 2000. PMID: 10908577 Free PMC article.
-
A Rev-CBP80-eIF4AI complex drives Gag synthesis from the HIV-1 unspliced mRNA.Nucleic Acids Res. 2018 Nov 30;46(21):11539-11552. doi: 10.1093/nar/gky851. Nucleic Acids Res. 2018. PMID: 30239828 Free PMC article.
-
A role for Rev in the association of HIV-1 gag mRNA with cytoskeletal beta-actin and viral protein expression.Biochimie. 1996;78(11-12):1075-80. doi: 10.1016/s0300-9084(97)86732-6. Biochimie. 1996. PMID: 9150887 Review.
-
Transcriptional and posttranscriptional regulation of HIV-1 gene expression.Cold Spring Harb Perspect Med. 2012 Feb;2(2):a006916. doi: 10.1101/cshperspect.a006916. Cold Spring Harb Perspect Med. 2012. PMID: 22355797 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

