Comparison of antiviral responses in two bat species reveals conserved and divergent innate immune pathways
- PMID: 37575178
- PMCID: PMC10415932
- DOI: 10.1016/j.isci.2023.107435
Comparison of antiviral responses in two bat species reveals conserved and divergent innate immune pathways
Abstract
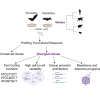
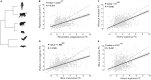
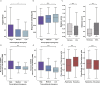
Bats host a range of disease-causing viruses without displaying clinical symptoms. The mechanisms behind this are a continuous source of interest. Here, we studied the antiviral response in the Egyptian fruit bat and Kuhl's pipistrelle, representing two subordinal clades. We profiled the antiviral response in fibroblasts using RNA sequencing and compared bat with primate and rodent responses. Both bats upregulate similar genes; however, a subset of these genes is transcriptionally divergent between them. These divergent genes also evolve rapidly in sequence, have specific promoter architectures, and are associated with programs underlying tolerance and resistance. Finally, we characterized antiviral genes that expanded in bats, with duplicates diverging in sequence and expression. Our study reveals a largely conserved antiviral program across bats and points to a set of genes that rapidly evolve through multiple mechanisms. These can contribute to bat adaptation to viral infection and provide directions to understanding the mechanisms behind it.
Keywords: Immunology; Phylogenetics; biological sciences; evolutionary biology; transcriptomics.
© 2023 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


References
LinkOut - more resources
Full Text Sources

