Identification of Immuno-Inflammation-Related Biomarkers for Acute Myocardial Infarction Based on Bioinformatics
- PMID: 37576155
- PMCID: PMC10417757
- DOI: 10.2147/JIR.S421196
Identification of Immuno-Inflammation-Related Biomarkers for Acute Myocardial Infarction Based on Bioinformatics
Abstract
Purpose: Previous studies have confirmed that inflammation and immunity are involved in the pathogenesis of acute myocardial infarction (AMI). However, only few related genes are identified as biomarkers for the diagnosis and treatment of AMI.
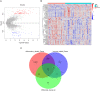
Patients and methods: GSE48060 and GSE60993 datasets were retrieved from Gene Expression Omnibus. The differentially expressed immuno-inflammation-related genes (DEIIRGs) were obtained from GSE48060, and the biomarkers for AMI were screened and validated using the "Neuralnet" package and GSE60993 dataset. Further, the biomarker-based nomogram was constructed, and miRNAs, transcription factors (TFs), and potential drugs targeting the biomarkers were explored. Furthermore, immune infiltration analysis was analyzed in AMI. Finally, the biomarkers were verified by assessing their mRNA levels using real-time quantitative PCR (RT-qPCR).
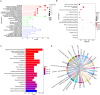
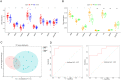
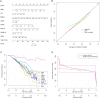
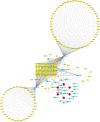
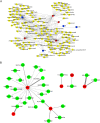
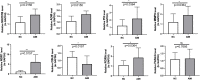
Results: First, eight biomarkers were screened via bioinformatics, and the artificial neural network model indicated a higher prediction accuracy for AMI even in the validation dataset. Nomogram had accurate forecasting ability for AMI as well. The TFs GTF2I, PHOX2B, RUNX1, and FOS targeting hsa-miR-1297 could regulate the expressions of ADM and CBLB, and RORA could effectively interact with melatonin and citalopram. RT-qPCR results for ADM, PI3, MMP9, NRG1 and CBLB were consistent with those of bioinformatic analysis.
Conclusion: In conclusion, eight key immuno-inflammation-related genes, namely, SH2D1B, ADM, PI3, MMP9, NRG1, CBLB, RORA, and FASLG, may serve as the potential biomarkers for AMI, in which the downregulation of CBLB and upregulation of ADM, PI3, and NRG1 in AMI was detected for the first time, providing a new strategy for the diagnosis and treatment of AMI.
Keywords: acute myocardial infarction; biomarker; immune-related gene; inflammation-related gene; nomogram.
© 2023 You and Dong.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflicts of interest.
Figures



References
-
- Anderson HVS, Masri SC, Abdallah MS, et al. 2022 ACC/AHA key data elements and definitions for chest pain and acute myocardial infarction: a report of the American Heart Association/American College of Cardiology Joint Committee on clinical data standards. J Am Coll Cardiol. 2022;80(17):1660–1700. doi: 10.1016/j.jacc.2022.05.012 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

