A high-throughput screening assay for silencing established HIV-1 macrophage infection identifies nucleoside analogs that perturb H3K9me3 on proviral genomes
- PMID: 37578230
- PMCID: PMC10506489
- DOI: 10.1128/jvi.00653-23
A high-throughput screening assay for silencing established HIV-1 macrophage infection identifies nucleoside analogs that perturb H3K9me3 on proviral genomes
Abstract
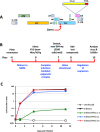
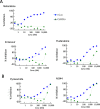
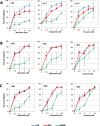
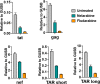
HIV-infected macrophages are long-lived cells that represent a barrier to functional cure. Additionally, low-level viral expression by central nervous system (CNS) macrophages contributes to neurocognitive deficits that develop despite antiretroviral therapy (ART). We recently identified H3K9me3 as an atypical epigenetic mark associated with chronic HIV infection in macrophages. Thus, strategies are needed to suppress HIV-1 expression in macrophages, but the unique myeloid environment and the responsible macrophage/CNS-tropic strains require cell/strain-specific approaches. Here, we generated an HIV-1 reporter virus from a CNS-derived strain with intact auxiliary genes expressing destabilized luciferase. We employed this reporter virus in polyclonal infection of primary human monocyte-derived macrophages (MDM) for a high-throughput screen (HTS) to identify compounds that suppress virus expression from established macrophage infection. Screening ~6,000 known drugs and compounds yielded 214 hits. A secondary screen with 10-dose titration identified 24 meeting criteria for HIV-selective activity. Using three replication-competent CNS-derived macrophage-tropic HIV-1 isolates and viral gene expression readout in MDM, we confirmed the effect of three purine analogs, nelarabine, fludarabine, and entecavir, showing the suppression of HIV-1 expression from established macrophage infection. Nelarabine inhibited the formation of H3K9me3 on HIV genomes in macrophages. Thus, this novel HTS assay can identify suppressors of HIV-1 transcription in established macrophage infection, such as nucleoside analogs and HDAC inhibitors, which may be linked to H3K9me3 modification. This screen may be useful to identify new metabolic and epigenetic agents that ameliorate HIV-driven neuroinflammation in people on ART or prevent viral recrudescence from macrophage reservoirs in strategies to achieve ART-free remission. IMPORTANCE Macrophages infected by HIV-1 are a long-lived reservoir and a barrier in current efforts to achieve HIV cure and also contribute to neurocognitive complications in people despite antiretroviral therapy (ART). Silencing HIV expression in these cells would be of great value, but the regulation of HIV-1 in macrophages differs from T cells. We developed a novel high-throughput screen for compounds that can silence established infection of primary macrophages, and identified agents that downregulate virus expression and alter provirus epigenetic profiles. The significance of this assay is the potential to identify new drugs that act in the unique macrophage environment on relevant viral strains, which may contribute to adjunctive treatment for HIV-associated neurocognitive disorders and/or prevent viral rebound in efforts to achieve ART-free remission or cure.
Keywords: HIV; epigenetics; high-throughput screen; macrophage; silencing.
Conflict of interest statement
Paul Lieberman declares his role as a founder of Vironika, LLC, as a Conflict of Interest that is managed by the Wistar Institute.
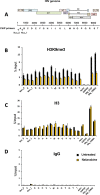
Figures

Similar articles
-
Epigenetic Suppression of HIV in Myeloid Cells by the BRD4-Selective Small Molecule Modulator ZL0580.J Virol. 2020 May 18;94(11):e01880-19. doi: 10.1128/JVI.01880-19. Print 2020 May 18. J Virol. 2020. PMID: 32188727 Free PMC article.
-
Epigenetic Landscape of HIV-1 Infection in Primary Human Macrophage.J Virol. 2022 Apr 13;96(7):e0016222. doi: 10.1128/jvi.00162-22. Epub 2022 Mar 23. J Virol. 2022. PMID: 35319230 Free PMC article.
-
Induction of Heme Oxygenase-1 Deficiency and Associated Glutamate-Mediated Neurotoxicity Is a Highly Conserved HIV Phenotype of Chronic Macrophage Infection That Is Resistant to Antiretroviral Therapy.J Virol. 2015 Oct;89(20):10656-67. doi: 10.1128/JVI.01495-15. Epub 2015 Aug 12. J Virol. 2015. PMID: 26269184 Free PMC article.
-
Brain macrophages harbor latent, infectious simian immunodeficiency virus.AIDS. 2019 Dec 1;33 Suppl 2(Suppl 2):S181-S188. doi: 10.1097/QAD.0000000000002269. AIDS. 2019. PMID: 31789817 Free PMC article. Review.
-
The HIV Reservoir in Monocytes and Macrophages.Front Immunol. 2019 Jun 26;10:1435. doi: 10.3389/fimmu.2019.01435. eCollection 2019. Front Immunol. 2019. PMID: 31297114 Free PMC article. Review.
Cited by
-
Investigating Antiprotozoal Chemotherapies with Novel Proteomic Tools-Chances and Limitations: A Critical Review.Int J Mol Sci. 2024 Jun 24;25(13):6903. doi: 10.3390/ijms25136903. Int J Mol Sci. 2024. PMID: 39000012 Free PMC article. Review.
-
Epigenetic modulation of myeloid cell functions in HIV and SARS-CoV-2 infection.Mol Biol Rep. 2024 Feb 24;51(1):342. doi: 10.1007/s11033-024-09266-2. Mol Biol Rep. 2024. PMID: 38400997 Free PMC article. Review.
References
-
- Collman R, Hassan NF, Walker R, Godfrey B, Cutilli J, Hastings JC, Friedman H, Douglas SD, Nathanson N. 1989. Infection of monocyte-derived macrophages with human immunodeficiency virus type 1 (HIV-1). monocyte-tropic and lymphocyte-tropic strains of HIV-1 show distinctive patterns of replication in a panel of cell types. J Exp Med 170:1149–1163. doi:10.1084/jem.170.4.1149 - DOI - PMC - PubMed
-
- Harris NS, Johnson AS, Huang Y-LA, Kern D, Fulton P, Smith DK, Valleroy LA, Hall HI. 2019. Vital signs: status of human immunodeficiency virus testing, viral suppression, and HIV preexposure prophylaxis - United States, 2013-2018. MMWR Morb Mortal Wkly Rep 68:1117–1123. doi:10.15585/mmwr.mm6848e1 - DOI - PMC - PubMed
-
- Métral M, Darling K, Locatelli I, Nadin I, Santos G, Brugger P, Kovari H, Cusini A, Gutbrod K, Tarr PE, Calmy A, Lecompte TD, Assal F, Monsch A, Kunze U, Stoeckle M, Schwind M, Schmid P, Pignatti R, Di Benedetto C, Du Pasquier R, Cavassini M, NAMACO study group, Swiss HIV Cohort Study . 2020. The Neurocognitive assessment in the metabolic and aging cohort (NAMACO) study: baseline participant profile. HIV Med 21:30–42. doi:10.1111/hiv.12795 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

