Quorum-sensing, intra- and inter-species competition in the staphylococci
- PMID: 37578829
- PMCID: PMC10482373
- DOI: 10.1099/mic.0.001381
Quorum-sensing, intra- and inter-species competition in the staphylococci
Abstract
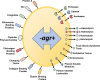
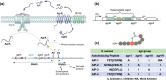
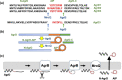
In Gram-positive bacteria such as Staphylococcus aureus and the coagulase-negative staphylococci (CoNS), the accessory gene regulator (agr) is a highly conserved but polymorphic quorum-sensing system involved in colonization, virulence and biofilm development. Signalling via agr depends on the interaction of an autoinducing peptide (AIP) with AgrC, a transmembrane sensor kinase that, once phosphorylated activates the response regulator AgrA. This in turn autoinduces AIP biosynthesis and drives target gene expression directly via AgrA or via the post-transcriptional regulator, RNAIII. In this review we describe the molecular mechanisms underlying the agr-mediated generation of, and response to, AIPs and the molecular basis of AIP-dependent activation and inhibition of AgrC. How the environment impacts on agr functionality is considered and the consequences of agr dysfunction for infection explored. We also discuss the concept of AIP-driven competitive interference between S. aureus and the CoNS and its anti-infective potential.
Keywords: Staphylococcus aureus; agr; autoinducing peptides; inter-bacterial competition; quorum sensing; staphylococci.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures
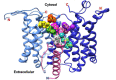
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

