Triple-negative breast tumors are dependent on mutant p53 for growth and survival
- PMID: 37579145
- PMCID: PMC10450424
- DOI: 10.1073/pnas.2308807120
Triple-negative breast tumors are dependent on mutant p53 for growth and survival
Abstract
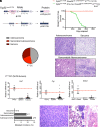
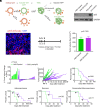
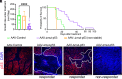
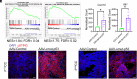
The TP53 tumor suppressor gene is mutated early in the majority of patients with triple-negative breast cancer (TNBC). The most frequent TP53 alterations are missense mutations that contribute to tumor aggressiveness. We developed an autochthonous somatic K14-Cre driven TNBC mouse model with p53R172H and p53R245W mutations in which mutant p53 can be toggled on and off genetically while leaving the tumor microenvironment intact and wild-type for p53. These mice develop TNBCs with a median latency of 1 y. Deletion of mutant p53R172H or p53R245W in vivo in these tumors blunts their tumor growth and significantly extends survival of mice. Downstream analyses revealed that deletion of mutant Trp53 activated the cyclic GMP-AMP Synthase-Stimulator of Interferon Genes pathway but did not cause apoptosis implicating other mechanisms of tumor regression. Furthermore, we determined that only tumors with stable mutant p53 are dependent on mutant p53 for growth.
Keywords: breast cancer; mouse models; mutant p53 stability; tumor cell vulnerabilities.
Conflict of interest statement
G.L. is on the scientific advisory board for PMV, receives honoraria from various universities for her lectures, and owns stock in PMV.
Figures


Similar articles
-
Mutant p53 protects triple-negative breast adenocarcinomas from ferroptosis in vivo.Sci Adv. 2024 Feb 16;10(7):eadk1835. doi: 10.1126/sciadv.adk1835. Epub 2024 Feb 14. Sci Adv. 2024. PMID: 38354236 Free PMC article.
-
p53R172H and p53R245W Hotspot Mutations Drive Distinct Transcriptomes in Mouse Mammary Tumors Through a Convergent Transcriptional Mediator.Cancer Res Commun. 2024 Aug 1;4(8):1991-2007. doi: 10.1158/2767-9764.CRC-24-0128. Cancer Res Commun. 2024. PMID: 38994678 Free PMC article.
-
Mutant p53 accumulation in human breast cancer is not an intrinsic property or dependent on structural or functional disruption but is regulated by exogenous stress and receptor status.J Pathol. 2014 Jul;233(3):238-46. doi: 10.1002/path.4356. Epub 2014 May 21. J Pathol. 2014. PMID: 24687952
-
P53 mutations in triple negative breast cancer upregulate endosomal recycling of epidermal growth factor receptor (EGFR) increasing its oncogenic potency.Crit Rev Oncol Hematol. 2013 Nov;88(2):284-92. doi: 10.1016/j.critrevonc.2013.05.003. Epub 2013 Jun 5. Crit Rev Oncol Hematol. 2013. PMID: 23755891 Review.
-
Targeting mutant p53: a key player in breast cancer pathogenesis and beyond.Cell Commun Signal. 2024 Oct 10;22(1):484. doi: 10.1186/s12964-024-01863-9. Cell Commun Signal. 2024. PMID: 39390510 Free PMC article. Review.
Cited by
-
CircCFL1 Promotes TNBC Stemness and Immunoescape via Deacetylation-Mediated c-Myc Deubiquitylation to Facilitate Mutant TP53 Transcription.Adv Sci (Weinh). 2024 Sep;11(34):e2404628. doi: 10.1002/advs.202404628. Epub 2024 Jul 9. Adv Sci (Weinh). 2024. PMID: 38981022 Free PMC article.
-
BUB1 Inhibition Sensitizes TNBC Cell Lines to Chemotherapy and Radiotherapy.Biomolecules. 2024 May 25;14(6):625. doi: 10.3390/biom14060625. Biomolecules. 2024. PMID: 38927028 Free PMC article.
-
Overcoming Chemoresistance in Cancer: The Promise of Crizotinib.Cancers (Basel). 2024 Jul 7;16(13):2479. doi: 10.3390/cancers16132479. Cancers (Basel). 2024. PMID: 39001541 Free PMC article. Review.
-
A cancer persistent DNA repair circuit driven by MDM2, MDM4 (MDMX), and mutant p53 for recruitment of MDC1 and 53BP1 on chromatin.Nucleic Acids Res. 2025 Jul 8;53(13):gkaf627. doi: 10.1093/nar/gkaf627. Nucleic Acids Res. 2025. PMID: 40626562 Free PMC article.
-
p53R245W Mutation Fuels Cancer Initiation and Metastases in NASH-driven Liver Tumorigenesis.Cancer Res Commun. 2023 Dec 29;3(12):2640-2652. doi: 10.1158/2767-9764.CRC-23-0218. Cancer Res Commun. 2023. PMID: 38047594 Free PMC article.
References
-
- Aschauer L., Muller P. A., Novel targets and interaction partners of mutant p53 Gain-Of-Function. Biochem. Soc. Trans. 44, 460–466 (2016). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

