Diversity-oriented synthesis encoded by deoxyoligonucleotides
- PMID: 37582753
- PMCID: PMC10427684
- DOI: 10.1038/s41467-023-40575-5
Diversity-oriented synthesis encoded by deoxyoligonucleotides
Erratum in
-
Author Correction: Diversity-oriented synthesis encoded by deoxyoligonucleotides.Nat Commun. 2023 Nov 21;14(1):7589. doi: 10.1038/s41467-023-43518-2. Nat Commun. 2023. PMID: 37989745 Free PMC article. No abstract available.
Abstract
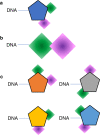
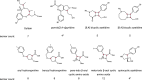
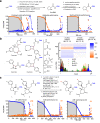
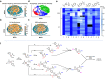
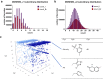
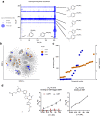
Diversity-oriented synthesis (DOS) is a powerful strategy to prepare molecules with underrepresented features in commercial screening collections, resulting in the elucidation of novel biological mechanisms. In parallel to the development of DOS, DNA-encoded libraries (DELs) have emerged as an effective, efficient screening strategy to identify protein binders. Despite recent advancements in this field, most DEL syntheses are limited by the presence of sensitive DNA-based constructs. Here, we describe the design, synthesis, and validation experiments performed for a 3.7 million-member DEL, generated using diverse skeleton architectures with varying exit vectors and derived from DOS, to achieve structural diversity beyond what is possible by varying appendages alone. We also show screening results for three diverse protein targets. We will make this DEL available to the academic scientific community to increase access to novel structural features and accelerate early-phase drug discovery.
© 2023. Springer Nature Limited.
Conflict of interest statement
C.J.G. is a shareholder in and employee of Kisbee Therapeutics. C.W.C. is an advisor to Anagenex. P.A.C. is an advisor to nference, Inc., Pfizer, Inc., and Belharra Therapeutics. B.M. is a scientific advisor to Magnet Biomedicine. S.L.S. is a shareholder and serves on the Board of Directors of Jnana Therapeutics and Kojin Therapeutics; is a shareholder and advises Kisbee Therapeutics, Belharra Therapeutics, Magnet Biomedicine, Exo Therapeutics, and Eikonizo Therapeutics; advises Vividian Therapeutics, Eisai Co., Ltd., Ono Pharma Foundation, F-Prime Capital Partners, and the Genomics Institute of the Novartis Research Foundation; and is a Novartis Faculty Scholar. All other authors declare no competing interests.
Figures
Similar articles
-
Diversity-oriented synthesis as a tool to expand the chemical space of DNA-encoded libraries.Bioorg Med Chem. 2021 Jul 1;41:116218. doi: 10.1016/j.bmc.2021.116218. Epub 2021 May 15. Bioorg Med Chem. 2021. PMID: 34030087 Review.
-
Strategies for developing DNA-encoded libraries beyond binding assays.Nat Chem. 2022 Feb;14(2):129-140. doi: 10.1038/s41557-021-00877-x. Epub 2022 Feb 4. Nat Chem. 2022. PMID: 35121833 Review.
-
Evolution of the Selection Methods of DNA-Encoded Chemical Libraries.Acc Chem Res. 2021 Sep 7;54(17):3491-3503. doi: 10.1021/acs.accounts.1c00375. Epub 2021 Aug 24. Acc Chem Res. 2021. PMID: 34427078
-
Recent Advances on the Selection Methods of DNA-Encoded Libraries.Chembiochem. 2021 Jul 15;22(14):2384-2397. doi: 10.1002/cbic.202100144. Epub 2021 May 7. Chembiochem. 2021. PMID: 33891355 Review.
-
N-Alkyl Linkers for DNA-Encoded Chemical Libraries.Chem Asian J. 2022 Apr 1;17(7):e202200016. doi: 10.1002/asia.202200016. Epub 2022 Mar 7. Chem Asian J. 2022. PMID: 35254005
Cited by
-
Is a Malleable Active Site Loop the Key to High Substrate Promiscuity? Hybrid, Biocatalytic Route to Structurally Diverse Taxoid Side Chains with Remarkable Dual Stereocontrol.Angew Chem Int Ed Engl. 2025 Jun 19:e202510889. doi: 10.1002/anie.202510889. Online ahead of print. Angew Chem Int Ed Engl. 2025. PMID: 40537411 Free PMC article.
-
Dataset Design for Building Models of Chemical Reactivity.ACS Cent Sci. 2023 Dec 8;9(12):2196-2204. doi: 10.1021/acscentsci.3c01163. eCollection 2023 Dec 27. ACS Cent Sci. 2023. PMID: 38161380 Free PMC article. Review.
-
Untargeted Diversity-Oriented Synthesis for the Discovery of New Antitumor Agents: An Integrated Approach of Inverse Virtual Screening, Bioinformatics, and Omics for Target Deconvolution.J Med Chem. 2025 Aug 14;68(15):16483-16517. doi: 10.1021/acs.jmedchem.5c01344. Epub 2025 Jul 24. J Med Chem. 2025. PMID: 40705771 Free PMC article.
-
Design, synthesis, and high-throughput in vitro anti-cancer evaluation of novel 4-aminopyrazolo[3,4-d]pyrimidine derivatives: potential anti-cancer candidates against UO-31 renal cancer cells.RSC Adv. 2024 Sep 27;14(42):30938-30953. doi: 10.1039/d4ra05136j. eCollection 2024 Sep 24. RSC Adv. 2024. PMID: 39346525 Free PMC article.
References
-
- Gerry CJ, Schreiber SL. Recent achievements and current trajectories of diversity-oriented synthesis. Curr. Opin. Chem. Biol. 2020;56:1–9. - PubMed
-
- Schreiber SL. Target-oriented and diversity-oriented organic synthesis in drug discovery. Science. 2000;287:1964–1969. - PubMed
-
- Galloway, W. R. J. D., Isidro-Llobet, A. & Spring, D. R. Diversity-oriented synthesis as a tool for the discovery of novel biologically active small molecules. Nat. Commun.1, 80 (2010). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

