AMBRA1 phosphorylation by CDK1 and PLK1 regulates mitotic spindle orientation
- PMID: 37584777
- PMCID: PMC10432340
- DOI: 10.1007/s00018-023-04878-6
AMBRA1 phosphorylation by CDK1 and PLK1 regulates mitotic spindle orientation
Abstract
AMBRA1 is a crucial factor for nervous system development, and its function has been mainly associated with autophagy. It has been also linked to cell proliferation control, through its ability to regulate c-Myc and D-type cyclins protein levels, thus regulating G1-S transition. However, it remains still unknown whether AMBRA1 is differentially regulated during the cell cycle, and if this pro-autophagy protein exerts a direct role in controlling mitosis too. Here we show that AMBRA1 is phosphorylated during mitosis on multiple sites by CDK1 and PLK1, two mitotic kinases. Moreover, we demonstrate that AMBRA1 phosphorylation at mitosis is required for a proper spindle function and orientation, driven by NUMA1 protein. Indeed, we show that the localization and/or dynamics of NUMA1 are strictly dependent on AMBRA1 presence, phosphorylation and binding ability. Since spindle orientation is critical for tissue morphogenesis and differentiation, our findings could account for an additional role of AMBRA1 in development and cancer ontogenesis.
Keywords: Cell cycle; Mitotic kinases; Mitotic spindle; NUMA1; Phosphorylation.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing financial interests.
Figures



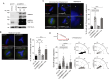


References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

