Model Organism Modifier (MOM): a user-friendly Galaxy workflow to detect modifiers from genome sequencing data using Caenorhabditis elegans
- PMID: 37585487
- PMCID: PMC10627290
- DOI: 10.1093/g3journal/jkad184
Model Organism Modifier (MOM): a user-friendly Galaxy workflow to detect modifiers from genome sequencing data using Caenorhabditis elegans
Abstract
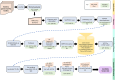
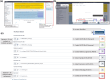
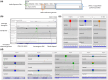
Genetic modifiers are variants modulating phenotypic outcomes of a primary detrimental variant. They contribute to rare diseases phenotypic variability, but their identification is challenging. Genetic screening with model organisms is a widely used method for demystifying genetic modifiers. Forward genetics screening followed by whole genome sequencing allows the detection of variants throughout the genome but typically produces thousands of candidate variants making the interpretation and prioritization process very time-consuming and tedious. Despite whole genome sequencing is more time and cost-efficient, usage of computational pipelines specific to modifier identification remains a challenge for biological-experiment-focused laboratories doing research with model organisms. To facilitate a broader implementation of whole genome sequencing in genetic screens, we have developed Model Organism Modifier or MOM, a pipeline as a user-friendly Galaxy workflow. Model Organism Modifier analyses raw short-read whole genome sequencing data and implements tailored filtering to provide a Candidate Variant List short enough to be further manually curated. We provide a detailed tutorial to run the Galaxy workflow Model Organism Modifier and guidelines to manually curate the Candidate Variant Lists. We have tested Model Organism Modifier on published and validated Caenorhabditis elegans modifiers screening datasets. As whole genome sequencing facilitates high-throughput identification of genetic modifiers in model organisms, Model Organism Modifier provides a user-friendly solution to implement the bioinformatics analysis of the short-read datasets in laboratories without expertise or support in Bioinformatics.
Keywords: Caenorhabditis elegans; bioinformatics pipeline; galaxy; genetic screening; modifiers; short-read whole genome sequencing.
© The Author(s) 2023. Published by Oxford University Press on behalf of The Genetics Society of America.
Conflict of interest statement
Conflicts of interest The author(s) declare no conflict of interest.
Figures

Similar articles
-
CloudMap: a cloud-based pipeline for analysis of mutant genome sequences.Genetics. 2012 Dec;192(4):1249-69. doi: 10.1534/genetics.112.144204. Epub 2012 Oct 10. Genetics. 2012. PMID: 23051646 Free PMC article.
-
Whole genome sequencing facilitates intragenic variant interpretation following modifier screening in C. elegans.BMC Genomics. 2021 Nov 13;22(1):820. doi: 10.1186/s12864-021-08142-8. BMC Genomics. 2021. PMID: 34773966 Free PMC article.
-
Mutation Mapping and Identification by Whole-Genome Sequencing.Methods Mol Biol. 2022;2468:257-269. doi: 10.1007/978-1-0716-2181-3_13. Methods Mol Biol. 2022. PMID: 35320569 Free PMC article.
-
Next-Generation Sequencing-Based Approaches for Mutation Mapping and Identification in Caenorhabditis elegans.Genetics. 2016 Oct;204(2):451-474. doi: 10.1534/genetics.115.186197. Genetics. 2016. PMID: 27729495 Free PMC article. Review.
-
Harnessing the power of genetics: fast forward genetics in Caenorhabditis elegans.Mol Genet Genomics. 2021 Jan;296(1):1-20. doi: 10.1007/s00438-020-01721-6. Epub 2020 Sep 4. Mol Genet Genomics. 2021. PMID: 32888055 Review.
Cited by
-
2024 VCP International Conference: Exploring multi-disciplinary approaches from basic science of valosin containing protein, an AAA+ ATPase protein, to the therapeutic advancement for VCP-associated multisystem proteinopathy.Neurobiol Dis. 2025 Apr;207:106861. doi: 10.1016/j.nbd.2025.106861. Epub 2025 Mar 2. Neurobiol Dis. 2025. PMID: 40037468 Review.
References
-
- Andrews S. FastQC: a quality control tool for high throughput sequence data. 2010.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
