Exploiting a living biobank to delineate mechanisms underlying disease-specific chromosome instability
- PMID: 37592171
- PMCID: PMC10435626
- DOI: 10.1007/s10577-023-09731-x
Exploiting a living biobank to delineate mechanisms underlying disease-specific chromosome instability
Abstract
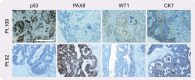
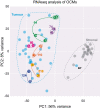
Chromosome instability (CIN) is a cancer hallmark that drives tumour heterogeneity, phenotypic adaptation, drug resistance and poor prognosis. High-grade serous ovarian cancer (HGSOC), one of the most chromosomally unstable tumour types, has a 5-year survival rate of only ~30% - largely due to late diagnosis and rapid development of drug resistance, e.g., via CIN-driven ABCB1 translocations. However, CIN is also a cell cycle vulnerability that can be exploited to specifically target tumour cells, illustrated by the success of PARP inhibitors to target homologous recombination deficiency (HRD). However, a lack of appropriate models with ongoing CIN has been a barrier to fully exploiting disease-specific CIN mechanisms. This barrier is now being overcome with the development of patient-derived cell cultures and organoids. In this review, we describe our progress building a Living Biobank of over 120 patient-derived ovarian cancer models (OCMs), predominantly from HGSOC. OCMs are highly purified tumour fractions with extensive proliferative potential that can be analysed at early passage. OCMs have diverse karyotypes, display intra- and inter-patient heterogeneity and mitotic abnormality rates far higher than established cell lines. OCMs encompass a broad-spectrum of HGSOC hallmarks, including a range of p53 alterations and BRCA1/2 mutations, and display drug resistance mechanisms seen in the clinic, e.g., ABCB1 translocations and BRCA2 reversion. OCMs are amenable to functional analysis, drug-sensitivity profiling, and multi-omics, including single-cell next-generation sequencing, and thus represent a platform for delineating HGSOC-specific CIN mechanisms. In turn, our vision is that this understanding will inform the design of new therapeutic strategies.
Keywords: Chromosome instability; cancer genomics; ovarian cancer; tumour heterogeneity.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





References
-
- Ahmed AA, Mills AD, Ibrahim AE, Temple J, Blenkiron C, Vias M, Massie CE, Iyer NG, McGeoch A, Crawford R, et al. The extracellular matrix protein TGFBI induces microtubule stabilization and sensitizes ovarian cancers to paclitaxel. Cancer Cell. 2007;12:514–527. doi: 10.1016/j.ccr.2007.11.014. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

