Analysis of tumor infiltrating CD4+ and CD8+ CDR3 sequences reveals shared features putatively associated to the anti-tumor immune response
- PMID: 37600765
- PMCID: PMC10436466
- DOI: 10.3389/fimmu.2023.1227766
Analysis of tumor infiltrating CD4+ and CD8+ CDR3 sequences reveals shared features putatively associated to the anti-tumor immune response
Abstract
Introduction: Tumor-infiltrating lymphocytes (TILs) have predictive and prognostic value in breast cancer (BC) and exert a protective function against tumor growth, indicating that it is susceptible to treatment using adoptive cell transfer of TILs or T cell receptor (TCR)-based therapies. TCR can be used to identify naturally tumor-reactive T cells, but little is known about the differences in the TCR repertoires of CD4+ and CD8+ TILs.
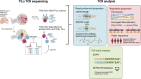
Methods: TCR high-throughput sequencing was performed using TILs derived from the initial cultures of 11 BC biopsies and expanded and sorted CD4+ and CD8+ TILs as well as using PBMCs from healthy donors expanded and sorted using the same methodology.
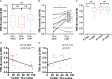
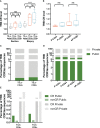
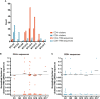
Results: Physicochemical TCR differences between T cell subsets were observed, as CD4+ TILs presented larger N(D)Nnt TRB sequences and with a higher usage of positively charged residues, although only the latest was also observed in peripheral T cells from healthy individuals. Moreover, in CD4+ TILs, a more restricted TCR repertoire with a higher abundance of similar sequences containing certain amino acid motifs was observed.
Discussion: Some differences between CD4+ and CD8+ TCRs were intrinsic to T cell subsets as can also be observed in peripheral T cells from healthy individuals, while other were only found in TILs samples and therefore may be tumor-driven. Notably, the higher similarity among CD4+ TCRs suggests a higher TCR promiscuity in this subset.
Keywords: CD4+ T cells; CD8+ T cells; T cell receptor; breast cancer; tumor-infiltrating lymphocytes.
Copyright © 2023 Aran, Lázaro, Marco, Molina, Abancó, Peg, Gión, Garrigós, Pérez-García, Cortés and Martí.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

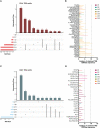

References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials

