The signature of cuproptosis-related immune genes predicts the tumor microenvironment and prognosis of prostate adenocarcinoma
- PMID: 37600770
- PMCID: PMC10433769
- DOI: 10.3389/fimmu.2023.1181370
The signature of cuproptosis-related immune genes predicts the tumor microenvironment and prognosis of prostate adenocarcinoma
Abstract
Background: Cuproptosis plays a crucial role in cancer, and different subtypes of cuproptosis have different immune profiles in prostate adenocarcinoma (PRAD). This study aimed to investigate immune genes associated with cuproptosis and develop a risk model to predict prognostic characteristics and chemotherapy/immunotherapy responses of patients with PRAD.
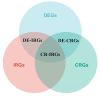
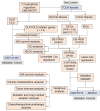
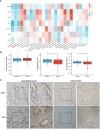
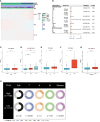
Methods: The CIBERSORT algorithm was used to evaluate the immune and stromal scores of patients with PRAD in The Cancer Genome Atlas (TCGA) cohort. Validation of differentially expressed genes DLAT and DLD in benign and malignant tissues by immunohistochemistry, and the immune-related genes of DLAT and DLD were further screened. Univariable Cox regression were performed to select key genes. Least absolute shrinkage and selection operator (LASSO)-Cox regression analyse was used to develop a risk model based on the selected genes. The model was validated in the TCGA, Memorial Sloan-Kettering Cancer Center (MSKCC) and Gene Expression Omnibus (GEO) datasets, as well as in this study unit cohort. The genes were examined via functional enrichment analysis, and the tumor immune features, tumor mutation features and copy number variations (CNVs) of patients with different risk scores were analysed. The response of patients to multiple chemotherapeutic/targeted drugs was assessed using the pRRophetic algorithm, and immunotherapy was inferred by the Tumor Immune Dysfunction and Exclusion (TIDE) and immunophenoscore (IPS).
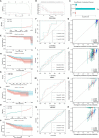
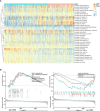
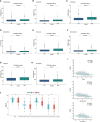
Results: Cuproptosis-related immune risk scores (CRIRSs) were developed based on PRLR, DES and LECT2. High CRIRSs indicated poor overall survival (OS), disease-free survival (DFS) in the TCGA-PRAD, MSKCC and GEO datasets and higher T stage and Gleason scores in TCGA-PRAD. Similarly, in the sample collected by the study unit, patients with high CRIRS had higher T-stage and Gleason scores. Additionally, higher CRIRSs were negatively correlated with the abundance of activated B cells, activated CD8+ T cells and other stromal or immune cells. The expression of some immune checkpoints was negatively correlated with CRIRSs. Tumor mutational burden (TMB), mutant-allele tumor heterogeneity (MATH) and copy number variation (CNV) scores were all higher in the high-CRIRS group. Multiple chemotherapeutic/targeted drugs and immunotherapy had better responsiveness in the low-CRIRS group.
Conclusion: Overall, lower CRIRS indicated better response to treatment strategies and better prognostic outcomes.
Keywords: LECT2; PrlR; cuproptosis; des; prostate cancer.
Copyright © 2023 Yao, Zhang, Li, Liu, Feng, Yan, Zhang and Xie.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures





References
-
- Lalonde E, Ishkanian AS, Sykes J, Fraser M, Ross-Adams H, Erho N, et al. Tumour genomic and microenvironmental heterogeneity for integrated prediction of 5-year biochemical recurrence of prostate cancer: a retrospective cohort study. Lancet Oncol (2014) 15(13):1521–32. doi: 10.1016/S1470-2045(14)71021-6 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

