Comparative analysis of gut DNA viromes in wild and captive Himalayan vultures
- PMID: 37601346
- PMCID: PMC10433386
- DOI: 10.3389/fmicb.2023.1120838
Comparative analysis of gut DNA viromes in wild and captive Himalayan vultures
Abstract
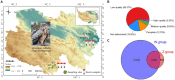
Introduction: Himalayan vultures (Gyps hinalayensis) are widely distributed on the Qinghai-Tibetan Plateau and play a crucial role in maintaining the ecological balance by feeding on decayed corpses of wild and domestic animals. Large-scale culture and metagenomics studies have broadened our understanding of viral diversity in animals' gastrointestinal tracts. However, despite the importance of gut viral communities in regulating bacterial diversity and performing symbiotic functions, no gut viral study has been conducted on Himalayan vultures. Furthermore, the impact of captivity on the gut virome of these vultures remains unknown.
Methods: In this study, metagenomic sequencing methods targeting DNA of virus-like particles enriched from feces were used to characterize the gut DNA viromes of wild and captive Himalayan vultures.
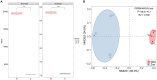
Results: In total, 22,938 unique viral operational taxonomic units (vOTUs) were identified and assigned to 140 viral genera in 41 viral families. These families included viruses associated with bacteria, animals, plants, insects, and archaea. Phage communities, including Siphoviridae, Microviridae, Myoviridae, Inoviridae, and Herelleviridae, dominated the gut virome of Himalayan vultures. Wild vultures exhibited higher viral richness and diversity compared with those in captivity. The functional capacity of the gut virome was characterized by identifying 93 KEGG pathways, which were significantly enriched in metabolism and genetic information processing. Abundant auxiliary metabolic genes, such as carbohydrate-active enzyme, and antibiotic resistance genes, were also found in the vultures' gut virome.
Discussion: Our findings reveal the complex and diverse viral community present in the gut virome of Himalayan vultures, which varies between wild, and captive states. The DNA virome dataset establishes a baseline for the vultures' gut virome and will serve as a reference for future virus isolation and cultivation. Understanding the impact of captivity on the gut virome contributes to our knowledge of vultures' response to captivity and aids in optimizing their rehabilitation and implementing protective measures.
Keywords: Gyps himalayensis; conservation biology; high-throughput sequencing technology; phage; scavenger; viral metagenomics; zoo.
Copyright © 2023 Zhai, Wang, Tang, Zheng, He, Zhao, Chen, Lin, Li, Bao, Lancuo, Sharshov, Liu and Wang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
A comparison of antibiotic resistance genes and mobile genetic elements in wild and captive Himalayan vultures.PeerJ. 2024 Jul 9;12:e17710. doi: 10.7717/peerj.17710. eCollection 2024. PeerJ. 2024. PMID: 39006014 Free PMC article.
-
Metagenomic comparison of gut communities between wild and captive Himalayan griffons.Front Vet Sci. 2024 May 9;11:1403932. doi: 10.3389/fvets.2024.1403932. eCollection 2024. Front Vet Sci. 2024. PMID: 38784654 Free PMC article.
-
Viral Metagenomic Analysis of the Fecal Samples in Domestic Dogs (Canis lupus familiaris).Viruses. 2023 Mar 6;15(3):685. doi: 10.3390/v15030685. Viruses. 2023. PMID: 36992396 Free PMC article.
-
The human gut virome: a multifaceted majority.Front Microbiol. 2015 Sep 11;6:918. doi: 10.3389/fmicb.2015.00918. eCollection 2015. Front Microbiol. 2015. PMID: 26441861 Free PMC article. Review.
-
A Review on Viral Metagenomics in Extreme Environments.Front Microbiol. 2019 Oct 18;10:2403. doi: 10.3389/fmicb.2019.02403. eCollection 2019. Front Microbiol. 2019. PMID: 31749771 Free PMC article. Review.
Cited by
-
The Potential of Co-Evolution and Interactions of Gut Bacteria-Phages in Bamboo-Eating Pandas: Insights from Dietary Preference-Based Metagenomic Analysis.Microorganisms. 2024 Mar 31;12(4):713. doi: 10.3390/microorganisms12040713. Microorganisms. 2024. PMID: 38674657 Free PMC article.
References
-
- Adawaren E. O., Mukandiwa L., Njoya E. M., Bekker L., Duncan N., Naidoo V. (2018). The use of liver slices from the cape vulture (Gyps coprotheres) to better understand the role of liver toxicity of non-steroidal anti-inflammatory drugs (NSAIDs) in vultures. Environ. Toxicol. Phar. 62, 147–155. doi: 10.1016/j.etap.2018.07.001, PMID: - DOI - PubMed
LinkOut - more resources
Full Text Sources

