Methods for the purification and detection of single nucleotide KRAS mutations on extrachromosomal circular DNA in human plasma
- PMID: 37602814
- PMCID: PMC10523981
- DOI: 10.1002/cam4.6385
Methods for the purification and detection of single nucleotide KRAS mutations on extrachromosomal circular DNA in human plasma
Abstract
Backgrounds: Despite recent advances, many cancers are still detected too late for curative treatment. There is, therefore, a need for the development of new diagnostic methods and biomarkers. One approach may arise from the detection of extrachromosomal circular DNA (eccDNA), which is part of cell-free DNA in human plasma.
Aims: First, we assessed and compared two methods for the purification of eccDNA from plasma. Second, we tested for an easy diagnostic application of eccDNA liquid biopsy-based assays.
Materials & methods: For the comparison we tested a solid-phase silica purification method and a phenol/chloroform method with salt precipitation. For the diagnostic application of eccDNA we developed and tested a qPCR primer-based SNP detection system, for the detection of two well-established cancer-causing KRAS mutations (G12V and G12R) on circular DNA. This investigation was supported by purifying, sequencing, and analysing clinical plasma samples for eccDNAs containing KRAS mutant alleles in 0.5 mL plasma from 16 pancreatic ductal adenocarcinoma patients and 19 healthy controls.
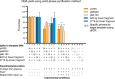
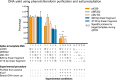
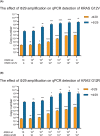
Results: In our method comparison we observed, that following exonuclease treatment a lower eccDNA yield was found for the phenol/chloroform method (15.7%-26.7%) compared with the solid-phase purification approach (47.8%-65.9%). For the diagnostic application of eccDNA tests, the sensitivity of the tested qPCR assay only reached ~10-3 in a background of 105 wild type (wt) KRAS circular entities, which was not improved by general amplification or primer-based inhibition of wt KRAS amplification. Furthermore, we did not detect eccDNA containing KRAS in any of the clinical samples.
Discussion: A potential explanation for our inability to detect any KRAS mutations in the clinical samples may be related to the general low abundance of eccDNA in plasma.
Conclusion: Taken together our results provide a benchmark for eccDNA purification methods while raising the question of what is required for the optimal fast and sensitive detection of SNP mutations on eccDNA with greater sensitivity than primer-based qPCR detection.
Keywords: KRAS mutations; circular DNA; eccDNA; liquid biopsy; phenol/chloroform DNA extraction; plasmids.
© 2023 The Authors. Cancer Medicine published by John Wiley & Sons Ltd.
Conflict of interest statement
LT and MM are employees of Roche. LBH, SFJ, JBN, SA, BR, and JSH declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


References
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

