Systematic Analysis of Diverse Polynucleotide Kinase Clp1 Family Proteins in Eukaryotes: Three Unique Clp1 Proteins of Trypanosoma brucei
- PMID: 37606665
- PMCID: PMC10598085
- DOI: 10.1007/s00239-023-10128-x
Systematic Analysis of Diverse Polynucleotide Kinase Clp1 Family Proteins in Eukaryotes: Three Unique Clp1 Proteins of Trypanosoma brucei
Abstract
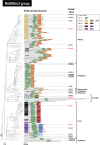
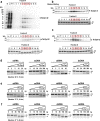
The Clp1 family proteins, consisting of the Clp1 and Nol9/Grc3 groups, have polynucleotide kinase (PNK) activity at the 5' end of RNA strands and are important enzymes in the processing of some precursor RNAs. However, it remains unclear how this enzyme family diversified in the eukaryotes. We performed a large-scale molecular evolutionary analysis of the full-length genomes of 358 eukaryotic species to classify the diverse Clp1 family proteins. The average number of Clp1 family proteins in eukaryotes was 2.3 ± 1.0, and most representative species had both Clp1 and Nol9/Grc3 proteins, suggesting that the Clp1 and Nol9/Grc3 groups were already formed in the eukaryotic ancestor by gene duplication. We also detected an average of 4.1 ± 0.4 Clp1 family proteins in members of the protist phylum Euglenozoa. For example, in Trypanosoma brucei, there are three genes of the Clp1 group and one gene of the Nol9/Grc3 group. In the Clp1 group proteins encoded by these three genes, the C-terminal domains have been replaced by unique characteristics domains, so we designated these proteins Tb-Clp1-t1, Tb-Clp1-t2, and Tb-Clp1-t3. Experimental validation showed that only Tb-Clp1-t2 has PNK activity against RNA strands. As in this example, N-terminal and C-terminal domain replacement also contributed to the diversification of the Clp1 family proteins in other eukaryotic species. Our analysis also revealed that the Clp1 family proteins in humans and plants diversified through isoforms created by alternative splicing.
Keywords: Family protein; Gene duplication; Molecular evolution; Polynucleotide kinase Clp1; Protein domain structure; RNA processing.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Adl SM, Bass D, Lane CE, Lukes J, Schoch CL, Smirnov A, Agatha S, Berney C, Brown MW, Burki F, Cardenas P, Cepicka I, Chistyakova L, Del Campo J, Dunthorn M, Edvardsen B, Eglit Y, Guillou L, Hampl V, Heiss AA, Hoppenrath M, James TY, Karnkowska A, Karpov S, Kim E, Kolisko M, Kudryavtsev A, Lahr DJG, Lara E, Le Gall L, Lynn DH, Mann DG, Massana R, Mitchell EAD, Morrow C, Park JS, Pawlowski JW, Powell MJ, Richter DJ, Rueckert S, Shadwick L, Shimano S, Spiegel FW, Torruella G, Youssef N, Zlatogursky V, Zhang Q. Revisions to the classification, nomenclature, and diversity of eukaryotes. J Eukaryot Microbiol. 2019;66:4. doi: 10.1111/jeu.12691. - DOI - PMC - PubMed
-
- Berthelot C, Brunet F, Chalopin D, Juanchich A, Bernard M, Noel B, Bento P, Da Silva C, Labadie K, Alberti A, Aury JM, Louis A, Dehais P, Bardou P, Montfort J, Klopp C, Cabau C, Gaspin C, Thorgaard GH, Boussaha M, Quillet E, Guyomard R, Galiana D, Bobe J, Volff JN, Genet C, Wincker P, Jaillon O, Roest Crollius H, Guiguen Y. The rainbow trout genome provides novel insights into evolution after whole-genome duplication in vertebrates. Nat Commun. 2014;5:3657. doi: 10.1038/ncomms4657. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

