Class I and II NADPH-cytochrome P450 reductases exhibit different roles in triterpenoid biosynthesis in Lotus japonicus
- PMID: 37621889
- PMCID: PMC10445947
- DOI: 10.3389/fpls.2023.1214602
Class I and II NADPH-cytochrome P450 reductases exhibit different roles in triterpenoid biosynthesis in Lotus japonicus
Abstract
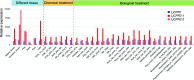
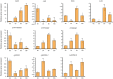
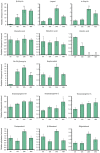
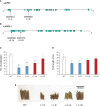
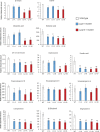
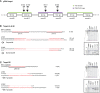
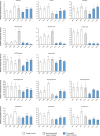
Cytochrome P450 monooxygenases (CYPs) are enzymes that play critical roles in the structural diversification of triterpenoids. To perform site-specific oxidations of the triterpene scaffold, CYPs require electrons transferred by NADPH-cytochrome P450 reductase (CPR), which is classified into two main classes, class I and class II, based on their structural difference. Lotus japonicus is a triterpenoids-producing model legume with one CPR class I gene (LjCPR1) and a minimum of two CPR class II genes (LjCPR2-1 and LjCPR2-2). CPR classes I and II from different plants have been reported to be involved in different metabolic pathways. By performing gene expression analyses of L. japonicus hairy root culture treated with methyl jasmonate (MeJA), this study revealed that LjCPR1, CYP716A51, and LUS were down-regulated which resulted in no change in betulinic acid and lupeol content. In contrast, LjCPR2s, bAS, CYP93E1, and CYP72A61 were significantly upregulated by MeJA treatment, followed by a significant increase of the precursors for soyasaponins, i.e. β-amyrin, 24-OH β-amyrin, and sophoradiol content. Triterpenoids profile analysis of LORE1 insertion and hairy root mutants showed that the loss of the Ljcpr2-1 gene significantly reduced soyasaponins precursors but not in Ljcpr1 mutants. However, Ljcpr1 and Ljcpr2-1 mutants showed a significant reduction in lupeol and oleanolic, ursolic, and betulinic acid contents. Furthermore, LjCPR1, but not LjCPR2, was crucial for seed development, supporting the previous notion that CPR class I might support plant basal metabolism. This study suggests that CPR classes I and II play different roles in L. japonicus triterpenoid biosynthesis.
Keywords: CRISPR/Cas9; LORE1; Lotus japonicus; NADPH-cytochrome P450 reductases (CPR); cytochrome P450 monooxygenases (CYP); triterpenoid biosynthesis.
Copyright © 2023 Istiandari, Yasumoto, Seki, Fukushima and Muranaka.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Lotus japonicus Triterpenoid Profile and Characterization of the CYP716A51 and LjCYP93E1 Genes Involved in Their Biosynthesis In Planta.Plant Cell Physiol. 2019 Nov 1;60(11):2496-2509. doi: 10.1093/pcp/pcz145. Plant Cell Physiol. 2019. PMID: 31418782
-
Comparative Analysis of NADPH-Cytochrome P450 Reductases From Legumes for Heterologous Production of Triterpenoids in Transgenic Saccharomyces cerevisiae.Front Plant Sci. 2021 Dec 16;12:762546. doi: 10.3389/fpls.2021.762546. eCollection 2021. Front Plant Sci. 2021. PMID: 34975947 Free PMC article.
-
Functional Characterization of CYP716 Family P450 Enzymes in Triterpenoid Biosynthesis in Tomato.Front Plant Sci. 2017 Jan 30;8:21. doi: 10.3389/fpls.2017.00021. eCollection 2017. Front Plant Sci. 2017. PMID: 28194155 Free PMC article.
-
Cytochrome P450 monooxygenase-mediated tailoring of triterpenoids and steroids in plants.Beilstein J Org Chem. 2022 Sep 21;18:1289-1310. doi: 10.3762/bjoc.18.135. eCollection 2022. Beilstein J Org Chem. 2022. PMID: 36225725 Free PMC article. Review.
-
Functional expression and regulation of eukaryotic cytochrome P450 enzymes in surrogate microbial cell factories.Eng Microbiol. 2022 Jan 19;2(1):100011. doi: 10.1016/j.engmic.2022.100011. eCollection 2022 Mar. Eng Microbiol. 2022. PMID: 39628612 Free PMC article. Review.
Cited by
-
Engineering Electron Transfer Pathway of Cytochrome P450s.Molecules. 2024 May 24;29(11):2480. doi: 10.3390/molecules29112480. Molecules. 2024. PMID: 38893355 Free PMC article. Review.
References
-
- DeLano W. L. (2020). The pyMOL molecular graphics system, version 2.3. Schrödinger LLC.
LinkOut - more resources
Full Text Sources
Research Materials

