Identifying immune cell infiltration and effective diagnostic biomarkers in Crohn's disease by bioinformatics analysis
- PMID: 37622114
- PMCID: PMC10445157
- DOI: 10.3389/fimmu.2023.1162473
Identifying immune cell infiltration and effective diagnostic biomarkers in Crohn's disease by bioinformatics analysis
Abstract
Background: Crohn's disease (CD) has an increasing incidence and prevalence worldwide. It is currently believed that both the onset and progression of the disease are closely related to immune system imbalance and the infiltration of immune cells. The aim of this study was to investigate the molecular immune mechanisms associated with CD and its fibrosis through bioinformatics analysis.
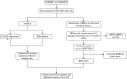
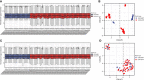
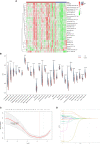
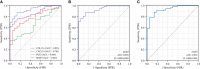
Methods: Three datasets from the Gene Expression Omnibus data base (GEO) were downloaded for data analysis and validation. Single sample gene enrichment analysis (ssGSEA) was used to evaluate the infiltration of immune cells in CD samples. Immune cell types with significant differences were identified by Wilcoxon test and Least Absolute Shrinkage and Selection Operator (LASSO) regression analysis. Differentially expressed genes (DEGs) were screened and then subjected to Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) functional correlation analysis, as well as protein-protein interaction (PPI) network analysis. The cytoHubba program and the GSE75214 dataset were used to screen for hub genes and plot Receiver operating characteristic (ROC)curves to screen for possible biomarkers of CD based on diagnostic efficacy. The hub genes of CD were correlated with five significantly different immune cells. In addition, validation was performed by real time quantitative PCR (RT-qPCR) experiments in colonic tissue of CD intestinal fibrosis rats to further identify hub genes that are more related to CD intestinal fibrosis.
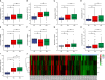
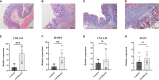
Results: The DEGs were analyzed separately by 10 algorithms and narrowed down to 9 DEGs after taking the intersection. 4 hub genes were further screened by the GSE75214 validation set, namely COL1A1, CXCL10, MMP2 and FGF2. COL1A1 has the highest specificity and sensitivity for the diagnosis of CD and is considered to have the potential to diagnose CD. Five immune cells with significant differences were screened between CD and health controls (HC). Through the correlation analysis between five kinds of immune cells and four biomarkers, it was found that CXCL10 was positively correlated with activated dendritic cells, effector memory CD8+ T cells. MMP2 was positively correlated with activated dendritic cells, gamma delta T cells (γδ T) and mast cells. MMP2 and COL1A1 were significantly increased in colon tissue of CD fibrosis rats.
Conclusion: MMP2, COL1A1, CXCL10 and FGF2 can be used as hub genes for CD. Among them, COL1A1 can be used as a biomarker for the diagnosis of CD. MMP2 and CXCL10 may be involved in the development and progression of CD by regulating activated dendritic cell, effector memory CD8+ T cell, γδ T cell and mast cell. In addition, MMP2 and COL1A1 may be more closely related to CD intestinal fibrosis.
Keywords: Crohn’s disease; biomarker; fibrosis; immune cells; ssGSEA.
Copyright © 2023 Huang, Wang, Chen, Chai, Qi, Zheng, Chen, Wu and Liu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




References
-
- Ng SC, Shi HY, Hamidi N, Underwood FE, Tang W, Benchimol EI, et al. Worldwide incidence and prevalence of inflammatory bowel disease in the 21st century: a systematic review of population-based studies. Lancet (London England). (2017) 390(10114):2769–78. doi: 10.1016/S0140-6736(17)32448-0 - DOI - PubMed
-
- Wu Kc, Liang J, Ran Zh, Qian JM, Yang H, Chen MH, et al. Consensus on diagnosis and treatment of inflammatory bowel disease (2018, Beijing). Chin J Digestion (2018) 38(5):292–311. doi: 10.19538/j.nk2018090106 - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

