Activation of a Vibrio cholerae CBASS anti-phage system by quorum sensing and folate depletion
- PMID: 37623317
- PMCID: PMC10653837
- DOI: 10.1128/mbio.00875-23
Activation of a Vibrio cholerae CBASS anti-phage system by quorum sensing and folate depletion
Abstract
To counteract infection with phage, bacteria have evolved a myriad of molecular defense systems. Some of these systems initiate a process called abortive infection, in which the infected cell kills itself to prevent phage propagation. However, such systems must be inhibited in the absence of phage infection to prevent spurious death of the host. Here, we show that the cyclic oligonucleotide based anti-phage signaling system (CBASS) accomplishes this by sensing intracellular folate molecules and only expressing this system in a group. These results enhance our understanding of the evolution of the seventh Vibrio cholerae pandemic and more broadly how bacteria defend themselves against phage infection.
Keywords: CD-NTases; cyclic GMP-AMP; folate; phage; quorum sensing.
Conflict of interest statement
The authors declare no conflict of interest.
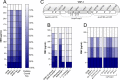
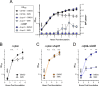
Figures



References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

