TMED10 mediates the loading of neosynthesised Sonic Hedgehog in COPII vesicles for efficient secretion and signalling
- PMID: 37624561
- PMCID: PMC11072717
- DOI: 10.1007/s00018-023-04918-1
TMED10 mediates the loading of neosynthesised Sonic Hedgehog in COPII vesicles for efficient secretion and signalling
Abstract
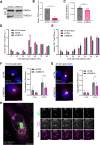
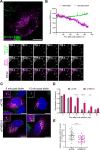
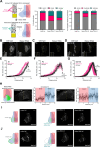
The morphogen Sonic Hedgehog (SHH) plays an important role in coordinating embryonic development. Short- and long-range SHH signalling occurs through a variety of membrane-associated and membrane-free forms. However, the molecular mechanisms that govern the early events of the trafficking of neosynthesised SHH in mammalian cells are still poorly understood. Here, we employed the retention using selective hooks (RUSH) system to show that newly-synthesised SHH is trafficked through the classical biosynthetic secretory pathway, using TMED10 as an endoplasmic reticulum (ER) cargo receptor for efficient ER-to-Golgi transport and Rab6 vesicles for Golgi-to-cell surface trafficking. TMED10 and SHH colocalized at ER exit sites (ERES), and TMED10 depletion significantly delays SHH loading onto ERES and subsequent exit leading to significant SHH release defects. Finally, we utilised the Drosophila wing imaginal disc model to demonstrate that the homologue of TMED10, Baiser (Bai), participates in Hedgehog (Hh) secretion and signalling in vivo. In conclusion, our work highlights the role of TMED10 in cargo-specific egress from the ER and sheds light on novel important partners of neosynthesised SHH secretion with potential impact on embryonic development.
Keywords: Biosynthetic pathway; ERES; Endoplasmic reticulum; Membrane trafficking; p24 proteins.
© 2023. The Author(s), under exclusive licence to Springer Nature Switzerland AG.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


References
-
- Wolpert L (2016). Positional information and pattern formation. In: Current topics in developmental biology. Vol. 117. Elsevier Inc. 10.1016/bs.ctdb.2015.11.008. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

