Mass Spectrometry Imaging of In Vitro Cryptosporidium parvum-Infected Cells and Host Tissue
- PMID: 37627264
- PMCID: PMC10452350
- DOI: 10.3390/biom13081200
Mass Spectrometry Imaging of In Vitro Cryptosporidium parvum-Infected Cells and Host Tissue
Abstract
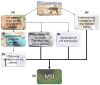
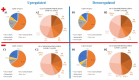
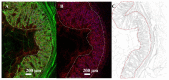
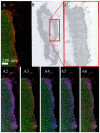
Cryptosporidium parvum is a zoonotic-relevant parasite belonging to the phylum Alveolata (subphylum Apicomplexa). One of the most zoonotic-relevant etiologies of cryptosporidiosis is the species C. parvum, infecting humans, cattle and wildlife. C. parvum-infected intestinal mucosa as well as host cells infected in vitro have not yet been the subject of extensive biochemical investigation. Efficient treatment options or vaccines against cryptosporidiosis are currently not available. Human cryptosporidiosis is currently known as a neglected poverty-related disease (PRD), being potentially fatal in young children or immunocompromised patients. In this study, we used a combination of atmospheric pressure scanning microprobe matrix-assisted laser desorption/ionization (AP-SMALDI) mass spectrometry imaging (MSI) and liquid chromatography-tandem mass spectrometry (HPLC-MS/MS) to determine and locate molecular biomarkers in in vitro C. parvum-infected host cells as well as parasitized neonatal calf intestines. Sections of C. parvum-infected and non-infected host cell pellets and infected intestines were examined to determine potential biomarkers. Human ileocecal adenocarcinoma cells (HCT-8) were used as a suitable in vitro host cell system. More than a thousand different molecular signals were found in both positive- and negative-ion mode, which were significantly increased in C. parvum-infected material. A database search in combination with HPLC-MS/MS experiments was employed for the structural verification of markers. Our results demonstrate some overlap between the identified markers and data obtained from earlier studies on other apicomplexan parasites. Statistically relevant biomarkers were imaged in cell layers of C. parvum-infected and non-infected host cells with 5 µm pixel size and in bovine intestinal tissue with 10 µm pixel size. This allowed us to substantiate their relevance once again. Taken together, the present approach delivers novel metabolic insights on neglected cryptosporidiosis affecting mainly children in developing countries.
Keywords: AP-SMALDI; Cryptosporidium parvum; mass spectrometry imaging.
Conflict of interest statement
B.S. is a consultant, S.G. an employee and N.H.A. is a part-time employee of TransMIT GmbH, Giessen, Germany. The other authors declare no conflict of interest.
Figures


References
-
- Minghui R., WHO . Ending the Neglect to Attain the Sustainable Development Goals: A Road Map for Neglected Tropical Diseases 2021−2030. WHO; Geneva, Switzerland: 2020. 196p
-
- Khalil I.A., Troeger C., Rao P.C., Blacker B.F., Brown A., Brewer T.G., Colombara D.V., De Hostos E.L., Engmann C., Guerrant R.L., et al. Morbidity, mortality, and long-term consequences associated with diarrhoea from Cryptosporidium infection in children younger than 5 years: A meta-analyses study. Lancet Glob. Health. 2018;6:E758–E768. doi: 10.1016/S2214-109X(18)30283-3. - DOI - PMC - PubMed
-
- Kotloff K.L., Nataro J.P., Blackwelder W.C., Nasrin D., Farag T.H., Panchalingam S., Wu Y., Sow S.O., Sur D., Breiman R.F., et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): A prospective, case-control study. Lancet. 2013;382:209–222. doi: 10.1016/S0140-6736(13)60844-2. - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

