Cellular Factors Involved in the Hepatitis D Virus Life Cycle
- PMID: 37632029
- PMCID: PMC10459925
- DOI: 10.3390/v15081687
Cellular Factors Involved in the Hepatitis D Virus Life Cycle
Abstract
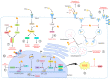
Hepatitis D virus (HDV) is a defective RNA virus with a negative-strand RNA genome encompassing less than 1700 nucleotides. The HDV genome encodes only for one protein, the hepatitis delta antigen (HDAg), which exists in two forms acting as nucleoproteins. HDV depends on the envelope proteins of the hepatitis B virus as a helper virus for packaging its ribonucleoprotein complex (RNP). HDV is considered the causative agent for the most severe form of viral hepatitis leading to liver fibrosis/cirrhosis and hepatocellular carcinoma. Many steps of the life cycle of HDV are still enigmatic. This review gives an overview of the complete life cycle of HDV and identifies gaps in knowledge. The focus is on the description of cellular factors being involved in the life cycle of HDV and the deregulation of cellular pathways by HDV with respect to their relevance for viral replication, morphogenesis and HDV-associated pathogenesis. Moreover, recent progress in antiviral strategies targeting cellular structures is summarized in this article.
Keywords: HBV; HDV; MVB; NTCP; RNA genome; morphogenesis; replication; surface protein.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
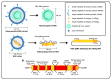
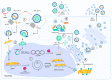
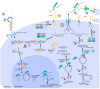
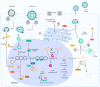
References
-
- Fattovich G., Giustina G., Christensen E., Pantalena M., Zagni I., Realdi G., Schalm S.W. Influence of hepatitis delta virus infection on morbidity and mortality in compensated cirrhosis type B. The European Concerted Action on Viral Hepatitis (Eurohep) Gut. 2000;46:420–426. doi: 10.1136/gut.46.3.420. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

