Neuronal ER-plasma membrane junctions couple excitation to Ca2+-activated PKA signaling
- PMID: 37633939
- PMCID: PMC10460453
- DOI: 10.1038/s41467-023-40930-6
Neuronal ER-plasma membrane junctions couple excitation to Ca2+-activated PKA signaling
Abstract
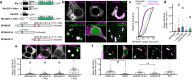
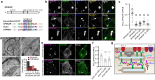
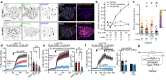
Junctions between the endoplasmic reticulum (ER) and the plasma membrane (PM) are specialized membrane contacts ubiquitous in eukaryotic cells. Concentration of intracellular signaling machinery near ER-PM junctions allows these domains to serve critical roles in lipid and Ca2+ signaling and homeostasis. Subcellular compartmentalization of protein kinase A (PKA) signaling also regulates essential cellular functions, however, no specific association between PKA and ER-PM junctional domains is known. Here, we show that in brain neurons type I PKA is directed to Kv2.1 channel-dependent ER-PM junctional domains via SPHKAP, a type I PKA-specific anchoring protein. SPHKAP association with type I PKA regulatory subunit RI and ER-resident VAP proteins results in the concentration of type I PKA between stacked ER cisternae associated with ER-PM junctions. This ER-associated PKA signalosome enables reciprocal regulation between PKA and Ca2+ signaling machinery to support Ca2+ influx and excitation-transcription coupling. These data reveal that neuronal ER-PM junctions support a receptor-independent form of PKA signaling driven by membrane depolarization and intracellular Ca2+, allowing conversion of information encoded in electrical signals into biochemical changes universally recognized throughout the cell.
© 2023. Springer Nature Limited.
Conflict of interest statement
The authors declare no competing interests.
Figures





References
-
- Saheki Y, De Camilli P. Endoplasmic reticulum-plasma membrane contact sites. Annu. Rev. Biochem. 2017;86:659–684. - PubMed
-
- Du J, Tao-Cheng JH, Zerfas P, McBain CJ. The K+ channel, Kv2.1, is apposed to astrocytic processes and is associated with inhibitory postsynaptic membranes in hippocampal and cortical principal neurons and inhibitory interneurons. Neuroscience. 1998;84:37–48. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

