MELLODDY: Cross-pharma Federated Learning at Unprecedented Scale Unlocks Benefits in QSAR without Compromising Proprietary Information
- PMID: 37642660
- PMCID: PMC11005050
- DOI: 10.1021/acs.jcim.3c00799
MELLODDY: Cross-pharma Federated Learning at Unprecedented Scale Unlocks Benefits in QSAR without Compromising Proprietary Information
Abstract
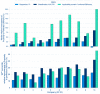
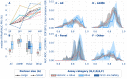
Federated multipartner machine learning has been touted as an appealing and efficient method to increase the effective training data volume and thereby the predictivity of models, particularly when the generation of training data is resource-intensive. In the landmark MELLODDY project, indeed, each of ten pharmaceutical companies realized aggregated improvements on its own classification or regression models through federated learning. To this end, they leveraged a novel implementation extending multitask learning across partners, on a platform audited for privacy and security. The experiments involved an unprecedented cross-pharma data set of 2.6+ billion confidential experimental activity data points, documenting 21+ million physical small molecules and 40+ thousand assays in on-target and secondary pharmacodynamics and pharmacokinetics. Appropriate complementary metrics were developed to evaluate the predictive performance in the federated setting. In addition to predictive performance increases in labeled space, the results point toward an extended applicability domain in federated learning. Increases in collective training data volume, including by means of auxiliary data resulting from single concentration high-throughput and imaging assays, continued to boost predictive performance, albeit with a saturating return. Markedly higher improvements were observed for the pharmacokinetics and safety panel assay-based task subsets.
Conflict of interest statement
The authors declare the following competing financial interest(s): M.N.G. and M.T. are employed and own stocks in the company Owkin commercializing the underlying Federated Learning Platform based on the open source Substra software. The remaining authors have no conflicts of interest to declare.
Figures




References
-
- Hansch C.; Maloney P. P.; Fujita T.; Muir R. M. Correlation of Biological Activity of Phenoxyacetic Acids with Hammett Substituent Constants and Partition Coefficients. Nature 1962, 194, 178–180. 10.1038/194178b0. - DOI
-
- Hansch C.; Fujita T. ρ-σ-π Analysis. A Method for the Correlation of Biological Activity and Chemical Structure. J. Am. Chem. Soc. 1964, 86, 1616–1626. 10.1021/ja01062a035. - DOI
-
- Muratov E. N.; Bajorath J.; Sheridan R. P.; Tetko I. V.; Filimonov D.; Poroikov V.; Oprea T. I.; Baskin I. I.; Varnek A.; Roitberg A.; Isayev O.; Curtalolo S.; Fourches D.; Cohen Y.; Aspuru-Guzik A.; Winkler D. A.; Agrafiotis D.; Cherkasov A.; Tropsha A. QSAR without Borders. Chem. Soc. Rev. 2020, 49, 3525–3564. 10.1039/D0CS00098A. - DOI - PMC - PubMed
-
- Tang Y.; Chen K. X.; Jiang H. L.; Ji R. Y. QSAR/QSTR of Fluoroquinolones: An Example of Simultaneous Analysis of Multiple Biological Activities Using Neural Network Method. Eur. J. Med. Chem. 1998, 33, 647–658. 10.1016/S0223-5234(98)80023-8. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

