Integrated multi-omics analysis reveals the molecular interplay between circadian clocks and cancer pathogenesis
- PMID: 37648722
- PMCID: PMC10469199
- DOI: 10.1038/s41598-023-39401-1
Integrated multi-omics analysis reveals the molecular interplay between circadian clocks and cancer pathogenesis
Abstract
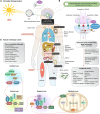
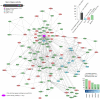
Circadian rhythms (CRs) are fundamental biological processes that significantly impact human well-being. Disruption of these rhythms can trigger insufficient neurocognitive development, insomnia, mental disorders, cardiovascular diseases, metabolic dysfunctions, and cancer. The field of chronobiology has increased our understanding of how rhythm disturbances contribute to cancer pathogenesis, and how circadian timing influences the efficacy of cancer treatments. As the circadian clock steadily gains recognition as an emerging factor in tumorigenesis, a thorough and comprehensive multi-omics analysis of CR genes/proteins has never been performed. To shed light on this, we performed, for the first time, an integrated data analysis encompassing genomic/transcriptomic alterations across 32 cancer types (n = 10,918 tumors) taken from the PanCancer Atlas, unfavorable prognostic protein analysis, protein-protein interactomics, and shortest distance score pathways to cancer hallmark phenotypes. This data mining strategy allowed us to unravel 31 essential CR-related proteins involved in the signaling crossroad between circadian rhythms and cancer. In the context of drugging the clock, we identified pharmacogenomic clinical annotations and drugs currently in late phase clinical trials that could be considered as potential cancer therapeutic strategies. These findings highlight the diverse roles of CR-related genes/proteins in the realm of cancer research and therapy.
© 2023. Springer Nature Limited.
Conflict of interest statement
The authors declare no competing interests.
Figures




References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials

