Use of adenine base editing and homology-independent targeted integration strategies to correct the cystic fibrosis causing variant, W1282X
- PMID: 37649273
- PMCID: PMC10656707
- DOI: 10.1093/hmg/ddad143
Use of adenine base editing and homology-independent targeted integration strategies to correct the cystic fibrosis causing variant, W1282X
Abstract
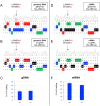
Small molecule drugs known as modulators can treat ~90% of people with cystic fibrosis (CF), but do not work for premature termination codon variants such as W1282X (c.3846G>A). Here we evaluated two gene editing strategies, Adenine Base Editing (ABE) to correct W1282X, and Homology-Independent Targeted Integration (HITI) of a CFTR superexon comprising exons 23-27 (SE23-27) to enable expression of a CFTR mRNA without W1282X. In Flp-In-293 cells stably expressing a CFTR expression minigene bearing W1282X, ABE corrected 24% of W1282X alleles, rescued CFTR mRNA from nonsense mediated decay and restored protein expression. However, bystander editing at the adjacent adenine (c.3847A>G), caused an amino acid change (R1283G) that affects CFTR maturation and ablates ion channel activity. In primary human nasal epithelial cells homozygous for W1282X, ABE corrected 27% of alleles, but with a notably lower level of bystander editing, and CFTR channel function was restored to 16% of wild-type levels. Using the HITI approach, correct integration of a SE23-27 in intron 22 of the CFTR locus in 16HBEge W1282X cells was detected in 5.8% of alleles, resulting in 7.8% of CFTR transcripts containing the SE23-27 sequence. Analysis of a clonal line homozygous for the HITI-SE23-27 produced full-length mature protein and restored CFTR anion channel activity to 10% of wild-type levels, which could be increased three-fold upon treatment with the triple combination of CF modulators. Overall, these data demonstrate two different editing strategies can successfully correct W1282X, the second most common class I variant, with a concomitant restoration of CFTR function.
Keywords: CFTR; CRISPR; HITI; W1282X; adenine base editing.
© The Author(s) 2023. Published by Oxford University Press.
Figures





Similar articles
-
Comparison of Cas9 and Cas12a CRISPR editing methods to correct the W1282X-CFTR mutation.J Cyst Fibros. 2022 Jan;21(1):181-187. doi: 10.1016/j.jcf.2021.05.014. Epub 2021 Jun 5. J Cyst Fibros. 2022. PMID: 34103250
-
Functional rescue of c.3846G>A (W1282X) in patient-derived nasal cultures achieved by inhibition of nonsense mediated decay and protein modulators with complementary mechanisms of action.J Cyst Fibros. 2020 Sep;19(5):717-727. doi: 10.1016/j.jcf.2019.12.001. Epub 2019 Dec 9. J Cyst Fibros. 2020. PMID: 31831337
-
Adenine base editing of CFTR using receptor targeted nanoparticles restores function to G542X cystic fibrosis airway epithelial cells.Cell Mol Life Sci. 2025 Apr 7;82(1):144. doi: 10.1007/s00018-025-05587-y. Cell Mol Life Sci. 2025. PMID: 40192756 Free PMC article.
-
Correction of Airway Stem Cells: Genome Editing Approaches for the Treatment of Cystic Fibrosis.Hum Gene Ther. 2020 Sep;31(17-18):956-972. doi: 10.1089/hum.2020.160. Epub 2020 Sep 8. Hum Gene Ther. 2020. PMID: 32741223 Free PMC article. Review.
-
Rewriting CFTR to cure cystic fibrosis.Prog Mol Biol Transl Sci. 2021;182:185-224. doi: 10.1016/bs.pmbts.2020.12.018. Epub 2021 Jan 28. Prog Mol Biol Transl Sci. 2021. PMID: 34175042 Review.
Cited by
-
Investigation of CFTR Function in Human Nasal Epithelial Cells Informs Personalized Medicine.Am J Respir Cell Mol Biol. 2024 Nov;71(5):577-588. doi: 10.1165/rcmb.2023-0398OC. Am J Respir Cell Mol Biol. 2024. PMID: 39012815 Free PMC article.
-
Systematic deletion of symmetrical CFTR exons reveals new therapeutic targets for exon skipping antisense oligonucleotides.NAR Mol Med. 2024 Nov 6;1(4):ugae017. doi: 10.1093/narmme/ugae017. eCollection 2024 Oct. NAR Mol Med. 2024. PMID: 39582793 Free PMC article.
-
Adenine base editing with engineered virus-like particles rescues the CFTR mutation G542X in patient-derived intestinal organoids.iScience. 2025 Feb 21;28(3):111979. doi: 10.1016/j.isci.2025.111979. eCollection 2025 Mar 21. iScience. 2025. PMID: 40144632 Free PMC article.
-
Functional rescue of F508del-CFTR through revertant mutations introduced by CRISPR base editing.Mol Ther. 2025 Mar 5;33(3):970-985. doi: 10.1016/j.ymthe.2025.01.011. Epub 2025 Jan 10. Mol Ther. 2025. PMID: 39797401
References
-
- Guo, J., Garratt, A. and Hill, A. (2022) Worldwide rates of diagnosis and effective treatment for cystic fibrosis. J Cyst Fibros, 21, 456–462. - PubMed
-
- Kerem, B., Rommens, J.M., Buchanan, J.A. et al. (1989) Identification of the cystic fibrosis gene: genetic analysis. Science, 245, 1073–1080. - PubMed
-
- Riordan, J.R., Rommens, J.M., Kerem, B. et al. (1989) Identification of the cystic fibrosis gene: cloning and characterization of complementary DNA. Science, 245, 1066–1073. - PubMed
-
- Castellani, C. (2013) CFTR2: how will it help care? Paediatr Respir Rev, 14S, 2–5. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

