Bioprospecting for polyesterase activity relevant for PET degradation in marine Enterobacterales isolates
- PMID: 37649797
- PMCID: PMC10462454
- DOI: 10.3934/microbiol.2023027
Bioprospecting for polyesterase activity relevant for PET degradation in marine Enterobacterales isolates
Abstract
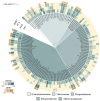
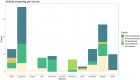
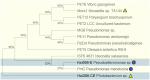
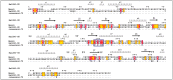
Plastics have quickly become an integral part of modern life. Due to excessive production and improper waste disposal, they are recognized as contaminants present in practically all habitat types. Although there are several polymers, polyethylene terephthalate (PET) is of particular concern due to its abundance in the environment. There is a need for a solution that is both cost-effective and ecologically friendly to address this pollutant. The use of microbial depolymerizing enzymes could offer a biological avenue for plastic degradation, though the full potential of these enzymes is yet to be uncovered. The purpose of this study was to use (1) plate-based screening methods to investigate the plastic degradation potential of marine bacteria from the order Enterobacterales collected from various organismal and environmental sources, and (2) perform genome-based analysis to identify polyesterases potentially related to PET degradation. 126 bacterial isolates were obtained from the strain collection of RD3, Research Unit Marine Symbioses-GEOMAR-and sequentially tested for esterase and polyesterase activity, in combination here referred to as PETase-like activity. The results show that members of the microbial families Alteromonadaceae, Shewanellaceae, and Vibrionaceae, derived from marine sponges and bryozoans, are the most promising candidates within the order Enterobacterales. Furthermore, 389 putative hydrolases from the α/β superfamily were identified in 23 analyzed genomes, of which 22 were sequenced for this study. Several candidates showed similarities with known PETases, indicating underlying enzymatic potential within the order Enterobacterales for PET degradation.
Keywords: PETase-like activity; bryozoans; marine bacteria; order Enterobacterales; sponges.
© 2023 the Author(s), licensee AIMS Press.
Conflict of interest statement
Conflict of interest: The authors declare no conflict of interest.
Figures


Similar articles
-
In silico Screening and Heterologous Expression of a Polyethylene Terephthalate Hydrolase (PETase)-Like Enzyme (SM14est) With Polycaprolactone (PCL)-Degrading Activity, From the Marine Sponge-Derived Strain Streptomyces sp. SM14.Front Microbiol. 2019 Oct 1;10:2187. doi: 10.3389/fmicb.2019.02187. eCollection 2019. Front Microbiol. 2019. PMID: 31632361 Free PMC article.
-
Identification of BgP, a Cutinase-Like Polyesterase From a Deep-Sea Sponge-Derived Actinobacterium.Front Microbiol. 2022 Apr 12;13:888343. doi: 10.3389/fmicb.2022.888343. eCollection 2022. Front Microbiol. 2022. PMID: 35495686 Free PMC article.
-
Using a marine microalga as a chassis for polyethylene terephthalate (PET) degradation.Microb Cell Fact. 2019 Oct 10;18(1):171. doi: 10.1186/s12934-019-1220-z. Microb Cell Fact. 2019. PMID: 31601227 Free PMC article.
-
Emerging Roles of PETase and MHETase in the Biodegradation of Plastic Wastes.Appl Biochem Biotechnol. 2021 Aug;193(8):2699-2716. doi: 10.1007/s12010-021-03562-4. Epub 2021 Apr 1. Appl Biochem Biotechnol. 2021. PMID: 33797026 Review.
-
Enzymatic Remediation of Polyethylene Terephthalate (PET)-Based Polymers for Effective Management of Plastic Wastes: An Overview.Front Bioeng Biotechnol. 2020 Nov 19;8:602325. doi: 10.3389/fbioe.2020.602325. eCollection 2020. Front Bioeng Biotechnol. 2020. PMID: 33330434 Free PMC article. Review.
Cited by
-
Ideonella sakaiensis Can Metabolize Bisphenol A as a Carbon Source.Microorganisms. 2023 Nov 30;11(12):2891. doi: 10.3390/microorganisms11122891. Microorganisms. 2023. PMID: 38138035 Free PMC article.
-
Linking the Metabolic Activity of Plastic-Degrading Fungi to Their Taxonomy and Evolution.J Fungi (Basel). 2025 May 15;11(5):378. doi: 10.3390/jof11050378. J Fungi (Basel). 2025. PMID: 40422712 Free PMC article. Review.
References
-
- Arnosti C, Bell C, Moorhead D, et al. Extracellular enzymes in terrestrial, freshwater, and marine environments: perspectives on system variability and common research needs. Biogeochemistry. 2014;117:5–21. doi: 10.1007/s10533-013-9906-5. - DOI
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
