Parkinson's disease alters the composition of subgingival microbiome
- PMID: 37649970
- PMCID: PMC10464550
- DOI: 10.1080/20002297.2023.2250650
Parkinson's disease alters the composition of subgingival microbiome
Abstract
Aim: The current study aimed to test the hypothesis that Parkinson's disease exacerbates periodontitis by altering its microbiome.
Materials and methods: Clinical periodontal parameters were recorded. Subgingival samples from healthy controls, periodontitis patients (PD), and Parkinson's patients with periodontitis (PA+PD) were analyzed using the checkerboard DNA-DNA hybridization technique for targeting 40 bacterial species typically associated with periodontal disease and health. Next-generation sequencing (NGS) of the 16S ribosomal RNA gene (V1-V3 regions) was performed to analyze the microbiome comprehensively.
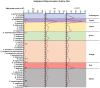
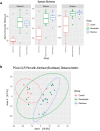
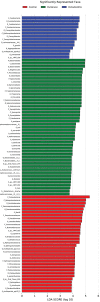
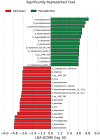
Results: Parkinson's patients had mild-to-moderate motor dysfunctions. Bleeding on probing was significantly increased in the PA+PD group compared to PD (p < 0.05). With checkerboard analysis, PA was associated with increased Treponema socranskii (p = 0.0062), Peptostreptococcaceae_[G-6] [Eubacterium]_nodatum (p = 0.0439), Parvimona micra (p < 0.0001), Prevotella melaninogenica (p = 0.0002), Lachnoanaerobaculum saburreum (p < 0.0001), and Streptococcus anginosus (p = 0.0020). Streptococcus intermedia (p = 0.0042), P.nodatum (p = 0.0022), P. micra (p = 0.0002), Treponema denticola (p = 0.0045), L.saburreum (p = 0.0267), P.melaninogenica (p = 0.0017), Campylobacter rectus (p = 0.0020), and T.socranskii (p = 0.0002) were higher; Aggregatibacter actinomycetemcomitans (p = 0.0072) was lower in deep pockets in the PA+PD compared to PD. Schaalia odontolytica (p = 0.0351) and A.actinomycetemcomitans (p = 0.002) were lower; C.rectus (p = 0.0002), P. micra (p = 0065), Streptococcus constellatus (p = 0.0151), T.denticola (p = 0.0141), P.melaninogenica (p = 0.0057), and T.socranskii (p = 0.0316) were higher in shallow pockets in the PA+PD. Diversity decreased in PD (p = 0.001) and PA+PD (p = 0.026) compared to control, with minimal differences in alpha and beta diversities among PD and PA+PD based on NGS results.
Conclusion: These data demonstrated that Parkinson's disease modifies PD-associated subgingival microbiome.
Keywords: Checkerboard; Parkinson’s disease; microbiome; microbiota; pathogenesis; periodontitis.
© 2023 The Author(s). Published by Informa UK Limited, trading as Taylor & Francis Group.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures





References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
