Small-Molecule Inhibition of KRAS through Conformational Selection
- PMID: 37663463
- PMCID: PMC10468774
- DOI: 10.1021/acsomega.3c04013
Small-Molecule Inhibition of KRAS through Conformational Selection
Abstract
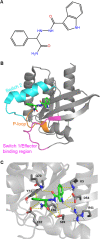
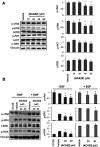
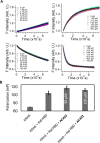
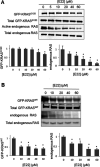
Mutations in KRAS account for about 20% of human cancers. Despite the major progress in recent years toward the development of KRAS inhibitors, including the discovery of covalent inhibitors of the G12C KRAS variant for the treatment of non-small-cell lung cancer, much work remains to be done to discover broad-acting inhibitors to treat many other KRAS-driven cancers. In a previous report, we showed that a 308.4 Da small-molecule ligand [(2R)-2-(N'-(1H-indole-3-carbonyl)hydrazino)-2-phenyl-acetamide] binds to KRAS with low micro-molar affinity [Chem. Biol. Drug Des.2019; 94(2):1441-1456]. Binding of this ligand, which we call ACA22, to the p1 pocket of KRAS and its interactions with residues at beta-strand 1 and the switch loops have been supported by data from nuclear magnetic resonance spectroscopy and microscale thermophoresis experiments. However, the inhibitory potential of the compound was not demonstrated. Here, we show that ACA22 inhibits KRAS-mediated signal transduction in cells expressing wild type (WT) and G12D mutant KRAS and reduces levels of guanosine triphosphate-loaded WT KRAS more effectively than G12D KRAS. We ruled out the direct effect on nucleotide exchange or effector binding as possible mechanisms of inhibition using a variety of biophysical assays. Combining these observations with binding data that show comparable affinities of the compound for the active and inactive forms of the mutant but not the WT, we propose conformational selection as a possible mechanism of action of ACA22.
© 2023 The Authors. Published by American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
Similar articles
-
A novel Imidazo[1,2-a]pyridine derivative modulates active KRASG12D through off-like conformational shifts in switch-I and switch-II regions, mimicking inactive KRASG12D.Int J Biol Macromol. 2024 Jun;270(Pt 2):132477. doi: 10.1016/j.ijbiomac.2024.132477. Epub 2024 May 19. Int J Biol Macromol. 2024. PMID: 38772459
-
KRAS Inhibitor that Simultaneously Inhibits Nucleotide Exchange Activity and Effector Engagement.ACS Bio Med Chem Au. 2022 Sep 26;2(6):617-626. doi: 10.1021/acsbiomedchemau.2c00045. eCollection 2022 Dec 21. ACS Bio Med Chem Au. 2022. PMID: 37101428 Free PMC article.
-
Modeling receptor flexibility in the structure-based design of KRASG12C inhibitors.J Comput Aided Mol Des. 2022 Aug;36(8):591-604. doi: 10.1007/s10822-022-00467-0. Epub 2022 Aug 5. J Comput Aided Mol Des. 2022. PMID: 35930206 Free PMC article.
-
The Therapeutic Landscape for KRAS-Mutated Colorectal Cancers.Cancers (Basel). 2023 Apr 19;15(8):2375. doi: 10.3390/cancers15082375. Cancers (Basel). 2023. PMID: 37190303 Free PMC article. Review.
-
Blockade of mutant RAS oncogenic signaling with a special emphasis on KRAS.Pharmacol Res. 2021 Oct;172:105806. doi: 10.1016/j.phrs.2021.105806. Epub 2021 Aug 24. Pharmacol Res. 2021. PMID: 34450320 Review.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

