Direct observation of tRNA-chaperoned folding of a dynamic mRNA ensemble
- PMID: 37673863
- PMCID: PMC10482949
- DOI: 10.1038/s41467-023-41155-3
Direct observation of tRNA-chaperoned folding of a dynamic mRNA ensemble
Abstract
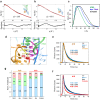
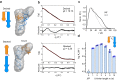
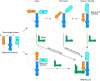
T-box riboswitches are multi-domain noncoding RNAs that surveil individual amino acid availabilities in most Gram-positive bacteria. T-boxes directly bind specific tRNAs, query their aminoacylation status to detect starvation, and feedback control the transcription or translation of downstream amino-acid metabolic genes. Most T-boxes rapidly recruit their cognate tRNA ligands through an intricate three-way stem I-stem II-tRNA interaction, whose establishment is not understood. Using single-molecule FRET, SAXS, and time-resolved fluorescence, we find that the free T-box RNA assumes a broad distribution of open, semi-open, and closed conformations that only slowly interconvert. tRNA directly binds all three conformers with distinct kinetics, triggers nearly instantaneous collapses of the open conformations, and returns the T-box RNA to their pre-binding conformations upon dissociation. This scissors-like dynamic behavior is enabled by a hinge-like pseudoknot domain which poises the T-box for rapid tRNA-induced domain closure. This study reveals tRNA-chaperoned folding of flexible, multi-domain mRNAs through a Venus flytrap-like mechanism.
© 2023. Springer Nature Limited.
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
Translational T-box riboswitches bind tRNA by modulating conformational flexibility.Nat Commun. 2024 Aug 3;15(1):6592. doi: 10.1038/s41467-024-50885-x. Nat Commun. 2024. PMID: 39097611 Free PMC article.
-
An evolving tale of two interacting RNAs-themes and variations of the T-box riboswitch mechanism.IUBMB Life. 2019 Aug;71(8):1167-1180. doi: 10.1002/iub.2098. Epub 2019 Jun 17. IUBMB Life. 2019. PMID: 31206978 Free PMC article. Review.
-
Structural and dynamic mechanisms for coupled folding and tRNA recognition of a translational T-box riboswitch.Nat Commun. 2023 Nov 15;14(1):7394. doi: 10.1038/s41467-023-43232-z. Nat Commun. 2023. PMID: 37968328 Free PMC article.
-
Structure and mechanism of the T-box riboswitches.Wiley Interdiscip Rev RNA. 2015 Jul-Aug;6(4):419-33. doi: 10.1002/wrna.1285. Epub 2015 May 8. Wiley Interdiscip Rev RNA. 2015. PMID: 25959893 Free PMC article. Review.
-
Unboxing the T-box riboswitches-A glimpse into multivalent and multimodal RNA-RNA interactions.Wiley Interdiscip Rev RNA. 2020 Nov;11(6):e1600. doi: 10.1002/wrna.1600. Epub 2020 Jul 6. Wiley Interdiscip Rev RNA. 2020. PMID: 32633085 Free PMC article. Review.
Cited by
-
RNA folding kinetics control riboswitch sensitivity in vivo.bioRxiv [Preprint]. 2024 Mar 29:2024.03.29.587317. doi: 10.1101/2024.03.29.587317. bioRxiv. 2024. Update in: Nat Commun. 2025 Jan 22;16(1):953. doi: 10.1038/s41467-024-55601-3. PMID: 38585885 Free PMC article. Updated. Preprint.
-
RNA folding kinetics control riboswitch sensitivity in vivo.Nat Commun. 2025 Jan 22;16(1):953. doi: 10.1038/s41467-024-55601-3. Nat Commun. 2025. PMID: 39843437 Free PMC article.
-
Structural basis of NEAT1 lncRNA maturation and menRNA instability.Nat Struct Mol Biol. 2024 Nov;31(11):1650-1654. doi: 10.1038/s41594-024-01361-z. Epub 2024 Jul 18. Nat Struct Mol Biol. 2024. PMID: 39026030
-
Translational T-box riboswitches bind tRNA by modulating conformational flexibility.Nat Commun. 2024 Aug 3;15(1):6592. doi: 10.1038/s41467-024-50885-x. Nat Commun. 2024. PMID: 39097611 Free PMC article.
-
Recognition of the tRNA structure: Everything everywhere but not all at once.Cell Chem Biol. 2024 Jan 18;31(1):36-52. doi: 10.1016/j.chembiol.2023.12.008. Epub 2023 Dec 29. Cell Chem Biol. 2024. PMID: 38159570 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

