Persister cell phenotypes contribute to poor patient outcomes after neoadjuvant chemotherapy in PDAC
- PMID: 37679568
- PMCID: PMC10518256
- DOI: 10.1038/s43018-023-00628-6
Persister cell phenotypes contribute to poor patient outcomes after neoadjuvant chemotherapy in PDAC
Abstract
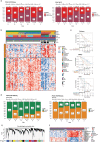
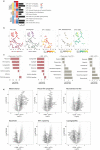
Neoadjuvant chemotherapy can improve the survival of individuals with borderline and unresectable pancreatic ductal adenocarcinoma; however, heterogeneous responses to chemotherapy remain a significant clinical challenge. Here, we performed RNA sequencing (n = 97) and multiplexed immunofluorescence (n = 122) on chemo-naive and postchemotherapy (post-CTX) resected patient samples (chemoradiotherapy excluded) to define the impact of neoadjuvant chemotherapy. Transcriptome analysis combined with high-resolution mapping of whole-tissue sections identified GATA6 (classical), KRT17 (basal-like) and cytochrome P450 3A (CYP3A) coexpressing cells that were preferentially enriched in post-CTX resected samples. The persistence of GATA6hi and KRT17hi cells post-CTX was significantly associated with poor survival after mFOLFIRINOX (mFFX), but not gemcitabine (GEM), treatment. Analysis of organoid models derived from chemo-naive and post-CTX samples demonstrated that CYP3A expression is a predictor of chemotherapy response and that CYP3A-expressing drug detoxification pathways can metabolize the prodrug irinotecan, a constituent of mFFX. These findings identify CYP3A-expressing drug-tolerant cell phenotypes in residual disease that may ultimately inform adjuvant treatment selection.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures















References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

