Integrative analysis of single-cell RNA-seq and ATAC-seq reveals heterogeneity of induced pluripotent stem cell-derived hepatic organoids
- PMID: 37680467
- PMCID: PMC10481365
- DOI: 10.1016/j.isci.2023.107675
Integrative analysis of single-cell RNA-seq and ATAC-seq reveals heterogeneity of induced pluripotent stem cell-derived hepatic organoids
Abstract
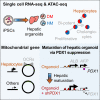
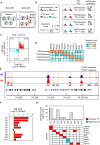
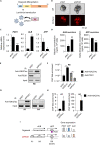
To gain deeper insights into transcriptomes and epigenomes of organoids, liver organoids from two states (expandable and more differentiated) were subjected to single-cell RNA-seq (scRNA-seq) and single-cell ATAC-seq (scATAC-seq) analyses. Mitochondrial gene expression was higher in differentiated than in non-differentiated hepatocytes, with ATAC-seq peaks increasing near the mitochondrial control region. Differentiation of liver organoids resulted in the expression of transcription factors that act as enhancers and repressors. In addition, epigenetic mechanisms regulating the expression of alpha-fetoprotein (AFP) and albumin (ALB) differed in liver organoids and adult liver. Knockdown of PDX1, an essential transcription factor for pancreas development, led to the hepatic maturation of liver organoids through regulation of AFP and ALB expression. This integrative analysis of the transcriptomes and epigenomes of liver organoids at the single-cell level may contribute to a better understanding of the regulatory networks during liver development and the further development of mature in vitro human liver models.
Keywords: Experimental models in systems biology; Integrative aspects of cell biology; Molecular mechanism of gene regulation; Transcriptomics.
© 2023 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
Generation of expandable human pluripotent stem cell-derived hepatocyte-like liver organoids.J Hepatol. 2019 Nov;71(5):970-985. doi: 10.1016/j.jhep.2019.06.030. Epub 2019 Jul 9. J Hepatol. 2019. PMID: 31299272
-
Generation of human liver organoids from pluripotent stem cell-derived hepatic endoderms.PeerJ. 2020 Oct 19;8:e9968. doi: 10.7717/peerj.9968. eCollection 2020. PeerJ. 2020. PMID: 33133779 Free PMC article.
-
Single-cell ATAC-seq of fetal human retina and stem-cell-derived retinal organoids shows changing chromatin landscapes during cell fate acquisition.Cell Rep. 2022 Jan 25;38(4):110294. doi: 10.1016/j.celrep.2021.110294. Cell Rep. 2022. PMID: 35081356
-
Single Cell Sequencing and Kidney Organoids Generated from Pluripotent Stem Cells.Clin J Am Soc Nephrol. 2020 Apr 7;15(4):550-556. doi: 10.2215/CJN.07470619. Epub 2020 Jan 28. Clin J Am Soc Nephrol. 2020. PMID: 31992574 Free PMC article. Review.
-
Single-cell RNA sequencing and kidney organoid differentiation.Clin Exp Nephrol. 2023 Jul;27(7):585-592. doi: 10.1007/s10157-023-02359-5. Epub 2023 May 20. Clin Exp Nephrol. 2023. PMID: 37209321 Free PMC article. Review.
Cited by
-
Guidelines for Manufacturing and Application of Organoids: Liver.Int J Stem Cells. 2024 May 30;17(2):120-129. doi: 10.15283/ijsc24044. Epub 2024 May 22. Int J Stem Cells. 2024. PMID: 38773747 Free PMC article.
-
Validating Well-Functioning Hepatic Organoids for Toxicity Evaluation.Toxics. 2024 May 17;12(5):371. doi: 10.3390/toxics12050371. Toxics. 2024. PMID: 38787150 Free PMC article.
-
Efficient and reproducible generation of human induced pluripotent stem cell-derived expandable liver organoids for disease modeling.Sci Rep. 2023 Dec 22;13(1):22935. doi: 10.1038/s41598-023-50250-w. Sci Rep. 2023. PMID: 38129682 Free PMC article.
-
Integrative genomic analyses reveal putative cell type-specific targets of the Drosophila ets transcription factor Pointed.BMC Genomics. 2024 Jan 23;25(1):103. doi: 10.1186/s12864-024-10017-7. BMC Genomics. 2024. PMID: 38262913 Free PMC article.
-
Utilization of Microfluidic Droplet-Based Methods in Diagnosis and Treatment Methods of Hepatocellular Carcinoma: A Review.Genes (Basel). 2024 Sep 25;15(10):1242. doi: 10.3390/genes15101242. Genes (Basel). 2024. PMID: 39457366 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

