FUS regulates the alternative splicing of cell proliferation genes related to atherosclerosis
- PMID: 37688506
- PMCID: PMC10666725
- DOI: 10.1177/15353702231187642
FUS regulates the alternative splicing of cell proliferation genes related to atherosclerosis
Abstract
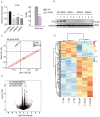
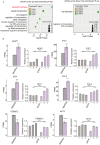
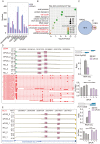
FUS plays a significant role as an RNA-binding protein in several cellular processes, including RNA splicing, DNA repair, and transcriptional regulation. However, the RNA-binding capacity of FUS in atherosclerosis is unclear. We aimed to study the functions of FUS in inflammatory regulation through the role of the splicing factor. We knocked down FUS with siRNA to further study the overall transcriptional level and select alternative splicing (AS) of FUS regulation in human umbilical vein endothelial cells (HUVECs) by RNA sequencing. The results suggested that the knockdown of FUS significantly affected gene expression in HUVECs. In addition, the knockdown of FUS resulted in 200 differentially expressed genes (DEGs) that were highly related to apoptotic process, signal transduction, multicellular organism development, cell adhesion and regulation of transcription, and DNA-templated pathways. Importantly, FUS extensively regulated 2870 AS events with a significant difference. Functional analysis of its modulated AS genes revealed they were highly enriched in cell cycle and cell population proliferation pathways. The qRT-PCR and RNA-seq data showed consistent results. Our findings suggested new knowledge of the mechanisms of FUS associated with atherosclerosis.
Keywords: FUS; RNA-binding protein; RNA-seq; alternative splicing; atherosclerosis.
Conflict of interest statement
Declaration of Conflicting InterestsThe author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Figures
Similar articles
-
Long noncoding RNA XXYLT1-AS2 regulates proliferation and adhesion by targeting the RNA binding protein FUS in HUVEC.Atherosclerosis. 2020 Apr;298:58-69. doi: 10.1016/j.atherosclerosis.2020.02.018. Epub 2020 Mar 4. Atherosclerosis. 2020. PMID: 32171981
-
Distinct and shared functions of ALS-associated proteins TDP-43, FUS and TAF15 revealed by multisystem analyses.Nat Commun. 2016 Jul 5;7:12143. doi: 10.1038/ncomms12143. Nat Commun. 2016. PMID: 27378374 Free PMC article.
-
HSPB1 Orchestrates the Inflammation-Associated Transcriptome Profile of Atherosclerosis in HUVECs.Front Biosci (Landmark Ed). 2025 Feb 17;30(2):36306. doi: 10.31083/FBL36306. Front Biosci (Landmark Ed). 2025. PMID: 40018940
-
FUS-mediated regulation of alternative RNA processing in neurons: insights from global transcriptome analysis.Wiley Interdiscip Rev RNA. 2016 May;7(3):330-40. doi: 10.1002/wrna.1338. Epub 2016 Jan 28. Wiley Interdiscip Rev RNA. 2016. PMID: 26822113 Review.
-
[The FUS protein: Physiological functions and a role in amyotrophic lateral sclerosis].Mol Biol (Mosk). 2017 May-Jun;51(3):387-399. doi: 10.7868/S0026898417020094. Mol Biol (Mosk). 2017. PMID: 28707655 Review. Russian.
Cited by
-
An integrative analysis of cell-specific transcriptomics and nuclear proteomics of sleep-deprived mouse cerebral cortex.bioRxiv [Preprint]. 2024 Sep 24:2024.09.24.611806. doi: 10.1101/2024.09.24.611806. bioRxiv. 2024. Update in: Sci Rep. 2025 Jul 29;15(1):27677. doi: 10.1038/s41598-025-10783-8. PMID: 39386443 Free PMC article. Updated. Preprint.
-
An integrative analysis of cell-specific transcriptomics and nuclear proteomics of sleep-deprived mouse cerebral cortex.Sci Rep. 2025 Jul 29;15(1):27677. doi: 10.1038/s41598-025-10783-8. Sci Rep. 2025. PMID: 40730573 Free PMC article.
-
The role of splicing events in the inflammatory response of atherosclerosis: molecular mechanisms and modulation.Front Immunol. 2024 Dec 17;15:1507420. doi: 10.3389/fimmu.2024.1507420. eCollection 2024. Front Immunol. 2024. PMID: 39742258 Free PMC article. Review.
References
-
- Falk E. Pathogenesis of atherosclerosis. J Am Coll Cardiol 2006;47:C7–12 - PubMed
-
- Peng J, Xiao X, Hu M, Zhang X. Interaction between gut microbiome and cardiovascular disease. Life Sci 2018;214:153–7 - PubMed
-
- Gimbrone MA, Jr, Topper JN, Nagel T, Anderson KR, Garcia-Cardeña G. Endothelial dysfunction, hemodynamic forces, and atherogenesis. Ann N Y Acad Sci 2000;902:230–9239 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

