Fluorescent Probes and Quenchers in Studies of Protein Folding and Protein-Lipid Interactions
- PMID: 37695081
- PMCID: PMC11113672
- DOI: 10.1002/tcr.202300232
Fluorescent Probes and Quenchers in Studies of Protein Folding and Protein-Lipid Interactions
Abstract
Fluorescence spectroscopy provides numerous methodological tools for structural and functional studies of biological macromolecules and their complexes. All fluorescence-based approaches require either existence of an intrinsic probe or an introduction of an extrinsic one. Moreover, studies of complex systems often require an additional introduction of a specific quencher molecule acting in combination with a fluorophore to provide structural or thermodynamic information. Here, we review the fundamentals and summarize the latest progress in applications of different classes of fluorescent probes and their specific quenchers, aimed at studies of protein folding and protein-membrane interactions. Specifically, we discuss various environment-sensitive dyes, FRET probes, probes for short-distance measurements, and several probe-quencher pairs for studies of membrane penetration of proteins and peptides. The goals of this review are: (a) to familiarize the readership with the general concept that complex biological systems often require both a probe and a quencher to decipher mechanistic details of functioning and (b) to provide example of the immediate applications of the described methods.
Keywords: FRET; environment-sensitive; intrinsic and extrinsic fluorescent probe; protein-membrane interactions; short-distance fluorescence quenching.
© 2023 The Chemical Society of Japan & Wiley-VCH GmbH.
Figures







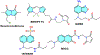



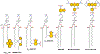
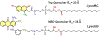

References
-
- Weber G, Annu. Rev. Biophys. Bioengin 1972, 1 (1) 553–570. - PubMed
-
- Yguerabide J, Foster MC, Fluorescence spectroscopy of biological membranes in Membrane spectroscopy, (Ed.: Grell E) Springer; Berlin Heidelberg: 1981, Vol. 31, pp 199–269. - PubMed
-
- Lakowicz JR, Principles of fluorescence spectroscopy. 3rd ed.; Springer: New York, 2006, p 954.
-
- Demchenko AP, Introduction to fluorescence sensing, Springer Science+Business Media B. V.: Netherlands, 2009; p 590.
-
- Ladokhin AS, Fluorescence spectroscopy in peptide and protein analysis in Encyclopedia of analytical chemistry: Applications of instrumental methods, John Wiley & Sons, Ltd, 2000, pp 5762–5779.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

