Diffuse midline glioma invasion and metastasis rely on cell-autonomous signaling
- PMID: 37702430
- PMCID: PMC10912010
- DOI: 10.1093/neuonc/noad161
Diffuse midline glioma invasion and metastasis rely on cell-autonomous signaling
Abstract
Background: Diffuse midline gliomas (DMG) are pediatric tumors with negligible 2-year survival after diagnosis characterized by their ability to infiltrate the central nervous system. In the hope of controlling the local growth and slowing the disease, all patients receive radiotherapy. However, distant progression occurs frequently in DMG patients. Current clues as to what causes tumor infiltration circle mainly around the tumor microenvironment, but there are currently no known determinants to predict the degree of invasiveness.
Methods: In this study, we use patient-derived glioma stem cells (GSCs) to create patient-specific 3D avatars to model interindividual invasion and elucidate the cellular supporting mechanisms.
Results: We show that GSC models in 3D mirror the invasive behavior of the parental tumors, thus proving the ability of DMG to infiltrate as an autonomous characteristic of tumor cells. Furthermore, we distinguished 2 modes of migration, mesenchymal and ameboid-like, and associated the ameboid-like modality with GSCs derived from the most invasive tumors. Using transcriptomics of both organoids and primary tumors, we further characterized the invasive ameboid-like tumors as oligodendrocyte progenitor-like, with highly contractile cytoskeleton and reduced adhesion ability driven by crucial over-expression of bone morphogenetic pathway 7 (BMP7). Finally, we deciphered MEK, ERK, and Rho/ROCK kinases activated downstream of the BMP7 stimulation as actionable targets controlling tumor cell motility.
Conclusions: Our findings identify 2 new therapeutic avenues. First, patient-derived GSCs represent a predictive tool for patient stratification in order to adapt irradiation strategies. Second, autocrine and short-range BMP7-related signaling becomes a druggable target to prevent DMG spread and metastasis.
Keywords: DMG-H3K27M; GSC; invasion; metastasis; tumor organoids.
© The Author(s) 2023. Published by Oxford University Press on behalf of the Society for Neuro-Oncology.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




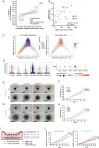


Comment in
-
Understanding the mechanisms of diffuse midline glioma cell migration toward therapeutic targeting.Neuro Oncol. 2024 Mar 4;26(3):569-570. doi: 10.1093/neuonc/noad263. Neuro Oncol. 2024. PMID: 38279953 Free PMC article. No abstract available.
References
-
- Schwartzentruber J, Korshunov A, Liu X-Y, et al. . Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature. 2012;482(7384):226–231. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

