A pan-sarcoma landscape of telomeric content shows that alterations in RAD51B and GID4 are associated with higher telomeric content
- PMID: 37709802
- PMCID: PMC10502097
- DOI: 10.1038/s41525-023-00369-6
A pan-sarcoma landscape of telomeric content shows that alterations in RAD51B and GID4 are associated with higher telomeric content
Abstract
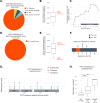
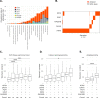
Tumor cells need to activate a telomere maintenance mechanism, enabling limitless replication. The bulk of evidence supports that sarcomas predominantly use alternative lengthening of telomeres (ALT) mechanism, commonly associated with alterations in ATRX and DAXX. In our dataset, only 12.3% of sarcomas harbored alterations in these genes. Thus, we checked for the presence of other genomic determinants of high telomeric content in sarcomas. Our dataset consisted of 13555 sarcoma samples, sequenced as a part of routine clinical care on the FoundationOne®Heme platform. We observed a median telomeric content of 622.3 telomeric reads per GC-matched million reads (TRPM) across all samples. In agreement with previous studies, telomeric content was significantly higher in ATRX altered and POT1 altered sarcomas. We further observed that sarcomas with alterations in RAD51B or GID4 were enriched in samples with high telomeric content, specifically within uterus leiomyosarcoma for RAD51B and soft tissue sarcoma (not otherwise specified, nos) for GID4, Furthermore, RAD51B and POT1 alterations were mutually exclusive with ATRX and DAXX alterations, suggestive of functional redundancy. Our results propose a role played by RAD51B and GID4 in telomere elongation in sarcomas and open research opportunities for agents aimed at targeting this critical pathway in tumorigenesis.
© 2023. Springer Nature Limited and Centre of Excellence in Genomic Medicine Research, King Abdulaziz University.
Conflict of interest statement
RS, DXJ, EE, GMF, LAA, and MM are employees of Foundation Medicine, Inc., a wholly owned subsidiary of Roche Holdings, Inc. and Roche Finance Ltd, and these employees have equity interest in an affiliate of these Roche entities. DT is the CEO of Omico, which provides national precision oncology services in Australia. DT has commercial and academic partnerships with multiple industry partners, including: Roche, Pfizer, Eisai, Astra Zeneca, Amgen, Abbvie, Eli Lilly, Beigene, Bayer, Illumina, Foundation Medicine, Microba, Merck, GMDx, Biotessellate, InterVenn. DT acts in a consultant capacity to the Health Futures Fund of Australian Unity, and the Maine Cancer Genome Initiative. All other authors don’t have conflicts of interest.
Figures



References
-
- Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–674. - PubMed
-
- Huffman KE, Levene SD, Tesmer VM, Shay JW, Wright WE. Telomere shortening is proportional to the size of the G-rich telomeric 3’-overhang. J. Biol. Chem. 2000;275:19719–19722. - PubMed
-
- Rahman R, Forsyth NR, Cui W. Telomeric 3′-overhang length is associated with the size of telomeres. Exp. Gerontol. 2008;43:258–265. - PubMed
-
- Martens UM, Chavez EA, Poon SS, Schmoor C, Lansdorp PM. Accumulation of short telomeres in human fibroblasts prior to replicative senescence. Exp. Cell Res. 2000;256:291–299. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

