Novel method of isolating nuclei of human oligodendrocyte precursor cells reveals substantial developmental changes in gene expression and H3K27ac histone modification
- PMID: 37712493
- PMCID: PMC10697634
- DOI: 10.1002/glia.24462
Novel method of isolating nuclei of human oligodendrocyte precursor cells reveals substantial developmental changes in gene expression and H3K27ac histone modification
Abstract
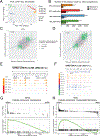
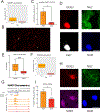
Oligodendrocyte precursor cells (OPCs) generate differentiated mature oligodendrocytes (MOs) during development. In adult brain, OPCs replenish MOs in adaptive plasticity, neurodegenerative disorders, and after trauma. The ability of OPCs to differentiate to MOs decreases with age and is compromised in disease. Here we explored the cell specific and age-dependent differences in gene expression and H3K27ac histone mark in these two cell types. H3K27ac is indicative of active promoters and enhancers. We developed a novel flow-cytometry-based approach to isolate OPC and MO nuclei from human postmortem brain and profiled gene expression and H3K27ac in adult and infant OPCs and MOs genome-wide. In adult brain, we detected extensive H3K27ac differences between the two cell types with high concordance between gene expression and epigenetic changes. Notably, the expression of genes that distinguish MOs from OPCs appears to be under a strong regulatory control by the H3K27ac modification in MOs but not in OPCs. Comparison of gene expression and H3K27ac between infants and adults uncovered numerous developmental changes in each cell type, which were linked to several biological processes, including cell proliferation and glutamate signaling. A striking example was a subset of histone genes that were highly active in infant samples but fully lost activity in adult brain. Our findings demonstrate a considerable rearrangement of the H3K27ac landscape that occurs during the differentiation of OPCs to MOs and during postnatal development of these cell types, which aligned with changes in gene expression. The uncovered regulatory changes justify further in-depth epigenetic studies of OPCs and MOs in development and disease.
Keywords: OPC; development; epigenetics.
© 2023 Wiley Periodicals LLC.
Conflict of interest statement
CONFLICT OF INTEREST STATEMENT
The authors declare that they have no conflict of interest.
Figures
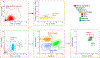




Similar articles
-
Chd7 Collaborates with Sox2 to Regulate Activation of Oligodendrocyte Precursor Cells after Spinal Cord Injury.J Neurosci. 2017 Oct 25;37(43):10290-10309. doi: 10.1523/JNEUROSCI.1109-17.2017. Epub 2017 Sep 20. J Neurosci. 2017. PMID: 28931573 Free PMC article.
-
Developmental trajectory of oligodendrocyte progenitor cells in the human brain revealed by single cell RNA sequencing.Glia. 2020 Jun;68(6):1291-1303. doi: 10.1002/glia.23777. Epub 2020 Jan 20. Glia. 2020. PMID: 31958186
-
Unbiased stereological analysis of the fate of oligodendrocyte progenitor cells in the adult mouse brain and effect of reference memory training.Behav Brain Res. 2017 Jun 30;329:127-139. doi: 10.1016/j.bbr.2017.04.027. Epub 2017 Apr 23. Behav Brain Res. 2017. PMID: 28442356
-
Brief review: Can modulating DNA methylation state help the clinical application of oligodendrocyte precursor cells as a source of stem cell therapy?Brain Res. 2019 Nov 15;1723:146386. doi: 10.1016/j.brainres.2019.146386. Epub 2019 Aug 13. Brain Res. 2019. PMID: 31419426 Free PMC article. Review.
-
From OPC to Oligodendrocyte: An Epigenetic Journey.Cells. 2019 Oct 11;8(10):1236. doi: 10.3390/cells8101236. Cells. 2019. PMID: 31614602 Free PMC article. Review.
References
-
- Barateiro A, Brites D and Fernandes A (2016) ‘Oligodendrocyte Development and Myelination in Neurodevelopment: Molecular Mechanisms in Health and Disease’, Current Pharmaceutical Design, 22, pp. 656–679. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

