Coarse-Grained Modeling Using Neural Networks Trained on Structural Data
- PMID: 37712507
- PMCID: PMC10569054
- DOI: 10.1021/acs.jctc.3c00516
Coarse-Grained Modeling Using Neural Networks Trained on Structural Data
Abstract
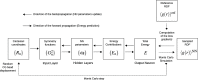
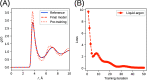
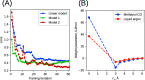
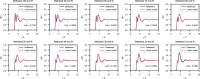
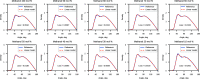
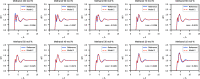
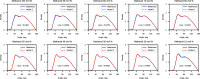
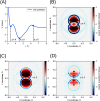
We propose a method of bottom-up coarse-graining, in which interactions within a coarse-grained model are determined by an artificial neural network trained on structural data obtained from multiple atomistic simulations. The method uses ideas of the inverse Monte Carlo approach, relating changes in the neural network weights with changes in average structural properties, such as radial distribution functions. As a proof of concept, we demonstrate the method on a system interacting by a Lennard-Jones potential modeled by a simple linear network and a single-site coarse-grained model of methanol-water solutions. In the latter case, we implement a nonlinear neural network with intermediate layers trained by atomistic simulations carried out at different methanol concentrations. We show that such a network acts as a transferable potential at the coarse-grained resolution for a wide range of methanol concentrations, including those not included in the training set.
Conflict of interest statement
The authors declare no competing financial interest.
Figures



Similar articles
-
Implicit solvent systematic coarse-graining of dioleoylphosphatidylethanolamine lipids: From the inverted hexagonal to the bilayer structure.PLoS One. 2019 Apr 5;14(4):e0214673. doi: 10.1371/journal.pone.0214673. eCollection 2019. PLoS One. 2019. PMID: 30951539 Free PMC article.
-
Thermodynamic Transferability in Coarse-Grained Force Fields Using Graph Neural Networks.J Chem Theory Comput. 2024 Dec 10;20(23):10524-10539. doi: 10.1021/acs.jctc.4c00788. Epub 2024 Nov 23. J Chem Theory Comput. 2024. PMID: 39579131 Free PMC article.
-
SAFT-γ force field for the simulation of molecular fluids. 1. A single-site coarse grained model of carbon dioxide.J Phys Chem B. 2011 Sep 29;115(38):11154-69. doi: 10.1021/jp204908d. Epub 2011 Sep 2. J Phys Chem B. 2011. PMID: 21815624
-
Bottom-Up Coarse-Grained Modeling of DNA.Front Mol Biosci. 2021 Mar 17;8:645527. doi: 10.3389/fmolb.2021.645527. eCollection 2021. Front Mol Biosci. 2021. PMID: 33816559 Free PMC article. Review.
-
Adaptive resolution simulations of biomolecular systems.Eur Biophys J. 2017 Dec;46(8):821-835. doi: 10.1007/s00249-017-1248-0. Epub 2017 Sep 13. Eur Biophys J. 2017. PMID: 28905203 Review.
Cited by
-
On the emergence of machine-learning methods in bottom-up coarse-graining.Curr Opin Struct Biol. 2025 Feb;90:102972. doi: 10.1016/j.sbi.2024.102972. Epub 2025 Jan 2. Curr Opin Struct Biol. 2025. PMID: 39752847 Review.
References
LinkOut - more resources
Full Text Sources
Research Materials

