Precision Oncology Comes of Age: Designing Best-in-Class Small Molecules by Integrating Two Decades of Advances in Chemistry, Target Biology, and Data Science
- PMID: 37712571
- PMCID: PMC10551669
- DOI: 10.1158/2159-8290.CD-23-0280
Precision Oncology Comes of Age: Designing Best-in-Class Small Molecules by Integrating Two Decades of Advances in Chemistry, Target Biology, and Data Science
Abstract
Small-molecule drugs have enabled the practice of precision oncology for genetically defined patient populations since the first approval of imatinib in 2001. Scientific and technology advances over this 20-year period have driven the evolution of cancer biology, medicinal chemistry, and data science. Collectively, these advances provide tools to more consistently design best-in-class small-molecule drugs against known, previously undruggable, and novel cancer targets. The integration of these tools and their customization in the hands of skilled drug hunters will be necessary to enable the discovery of transformational therapies for patients across a wider spectrum of cancers.
Significance: Target-centric small-molecule drug discovery necessitates the consideration of multiple approaches to identify chemical matter that can be optimized into drug candidates. To do this successfully and consistently, drug hunters require a comprehensive toolbox to avoid following the "law of instrument" or Maslow's hammer concept where only one tool is applied regardless of the requirements of the task. Combining our ever-increasing understanding of cancer and cancer targets with the technological advances in drug discovery described below will accelerate the next generation of small-molecule drugs in oncology.
©2023 The Authors; Published by the American Association for Cancer Research.
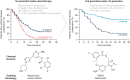
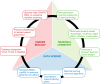
Figures



Similar articles
-
The FDA Oncology Center of Excellence and precision medicine.Exp Biol Med (Maywood). 2018 Feb;243(3):308-312. doi: 10.1177/1535370217740861. Epub 2017 Nov 6. Exp Biol Med (Maywood). 2018. PMID: 29105511 Free PMC article. Review.
-
Realizing Cancer Precision Medicine by Integrating Systems Biology and Nanomaterial Engineering.Adv Mater. 2020 Sep;32(35):e1906783. doi: 10.1002/adma.201906783. Epub 2020 Apr 6. Adv Mater. 2020. PMID: 32253807 Review.
-
Individualized network-based drug repositioning infrastructure for precision oncology in the panomics era.Brief Bioinform. 2017 Jul 1;18(4):682-697. doi: 10.1093/bib/bbw051. Brief Bioinform. 2017. PMID: 27296652 Review.
-
Top advances of the year: Precision oncology.Cancer. 2023 Jun 1;129(11):1634-1642. doi: 10.1002/cncr.34743. Epub 2023 Mar 22. Cancer. 2023. PMID: 36946766 Review.
-
Molecular tests and target therapies in oncology: recommendations from the Italian workshop.Future Oncol. 2021 Sep;17(26):3529-3539. doi: 10.2217/fon-2021-0286. Epub 2021 Jul 13. Future Oncol. 2021. PMID: 34254524
Cited by
-
Cancer epigenetic therapy: recent advances, challenges, and emerging opportunities.Epigenomics. 2025 Jan;17(1):59-74. doi: 10.1080/17501911.2024.2430169. Epub 2024 Nov 27. Epigenomics. 2025. PMID: 39601374 Free PMC article. Review.
-
From dominance to decline: can we reverse the trend in small molecule and TKI cancer therapies?Pathologica. 2025 Jun;117(3):199-203. doi: 10.32074/1591-951X-N869. Epub 2025 May 21. Pathologica. 2025. PMID: 40396419 Free PMC article. Review.
-
Self-Assembled Nanocomposite DOX/TPOR4@CB[7]4 for Enhanced Synergistic Photodynamic Therapy and Chemotherapy in Neuroblastoma.Pharmaceutics. 2024 Jun 18;16(6):822. doi: 10.3390/pharmaceutics16060822. Pharmaceutics. 2024. PMID: 38931942 Free PMC article.
-
Insights into the structural and functional activities of forgotten Kinases: PCTAIREs CDKs.Mol Cancer. 2024 Jun 29;23(1):135. doi: 10.1186/s12943-024-02043-6. Mol Cancer. 2024. PMID: 38951876 Free PMC article. Review.
-
Cyclin-dependent protein kinases and cell cycle regulation in biology and disease.Signal Transduct Target Ther. 2025 Jan 13;10(1):11. doi: 10.1038/s41392-024-02080-z. Signal Transduct Target Ther. 2025. PMID: 39800748 Free PMC article. Review.
References
-
- Smith DA, Di L, Kerns EH. The effect of plasma protein binding on in vivo efficacy: misconceptions in drug discovery. Nat Rev Drug Discov 2010;9:929–39. - PubMed
-
- Yap TA, Sandhu SK, Workman P, de Bono JS. Envisioning the future of early anticancer drug development. Nat Rev Cancer 2010;10:514–23. - PubMed
-
- Morgan P, Van Der Graaf PH, Arrowsmith J, Feltner DE, Drummond KS, Wegner CD, et al. Can the flow of medicines be improved? Fundamental pharmacokinetic and pharmacological principles toward improving phase II survival. Drug Discov Today 2012;17:419–24. - PubMed
-
- Haslam A, Kim MS, Prasad V. Updated estimates of eligibility for and response to genome-targeted oncology drugs among US cancer patients, 2006–2020. Ann Oncol 2021;32:926–32. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

