Rep protein accommodates together dsDNA and ssDNA which enables a loop-back mechanism to plasmid DNA replication initiation
- PMID: 37713613
- PMCID: PMC10602881
- DOI: 10.1093/nar/gkad740
Rep protein accommodates together dsDNA and ssDNA which enables a loop-back mechanism to plasmid DNA replication initiation
Abstract
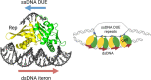
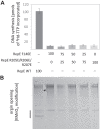
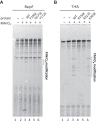
For DNA replication initiation in Bacteria, replication initiation proteins bind to double-stranded DNA (dsDNA) and interact with single-stranded DNA (ssDNA) at the replication origin. The structural-functional relationship of the nucleoprotein complex involving initiator proteins is still elusive and different models are proposed. In this work, based on crosslinking combined with mass spectrometry (MS), the analysis of mutant proteins and crystal structures, we defined amino acid residues essential for the interaction between plasmid Rep proteins, TrfA and RepE, and ssDNA. This interaction and Rep binding to dsDNA could not be provided in trans, and both are important for dsDNA melting at DNA unwinding element (DUE). We solved two crystal structures of RepE: one in a complex with ssDNA DUE, and another with both ssDNA DUE and dsDNA containing RepE-specific binding sites (iterons). The amino acid residues involved in interaction with ssDNA are located in the WH1 domain in stand β1, helices α1 and α2 and in the WH2 domain in loops preceding strands β1' and β2' and in these strands. It is on the opposite side compared to RepE dsDNA-recognition interface. Our data provide evidence for a loop-back mechanism through which the plasmid replication initiator molecule accommodates together dsDNA and ssDNA.
© The Author(s) 2023. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures






Similar articles
-
Sequence-specific interactions of Rep proteins with ssDNA in the AT-rich region of the plasmid replication origin.Nucleic Acids Res. 2014 Jul;42(12):7807-18. doi: 10.1093/nar/gku453. Epub 2014 May 16. Nucleic Acids Res. 2014. PMID: 24838560 Free PMC article.
-
Regulation of DNA replication by iterons: an interaction between the ori2 and incC regions mediated by RepE-bound iterons inhibits DNA replication of mini-F plasmid in Escherichia coli.EMBO J. 1999 Jul 1;18(13):3856-67. doi: 10.1093/emboj/18.13.3856. EMBO J. 1999. PMID: 10393200 Free PMC article.
-
The RepE initiator is a double-stranded and single-stranded DNA-binding protein that forms an atypical open complex at the onset of replication of plasmid pAMbeta 1 from Gram-positive bacteria.J Biol Chem. 2001 Mar 30;276(13):10234-46. doi: 10.1074/jbc.M010118200. Epub 2000 Dec 20. J Biol Chem. 2001. PMID: 11124267
-
Twenty years of the pPS10 replicon: insights on the molecular mechanism for the activation of DNA replication in iteron-containing bacterial plasmids.Plasmid. 2004 Sep;52(2):69-83. doi: 10.1016/j.plasmid.2004.06.002. Plasmid. 2004. PMID: 15336485 Review.
-
Two replication initiators - one mechanism for replication origin opening?Plasmid. 2014 Nov;76:72-8. doi: 10.1016/j.plasmid.2014.10.003. Epub 2014 Oct 18. Plasmid. 2014. PMID: 25454070 Review.
Cited by
-
Chlamydia plasmid-encoded protein Pgp2 is a replication initiator with a unique β-hairpin necessary for iteron-binding and plasmid replication.bioRxiv [Preprint]. 2024 Nov 15:2024.11.14.623704. doi: 10.1101/2024.11.14.623704. bioRxiv. 2024. Update in: Infect Immun. 2025 Mar 11;93(3):e0060224. doi: 10.1128/iai.00602-24. PMID: 39569140 Free PMC article. Updated. Preprint.
-
Chlamydia plasmid-encoded protein Pgp2 is a replication initiator with a unique β-hairpin necessary for iteron-binding and plasmid replication.Infect Immun. 2025 Mar 11;93(3):e0060224. doi: 10.1128/iai.00602-24. Epub 2025 Feb 7. Infect Immun. 2025. PMID: 39918305 Free PMC article.
-
Toward an understanding of the DNA replication initiation in bacteria.Front Microbiol. 2024 Jan 5;14:1328842. doi: 10.3389/fmicb.2023.1328842. eCollection 2023. Front Microbiol. 2024. PMID: 38249469 Free PMC article. Review.
References
-
- Bleichert F., Botchan M.R., Berger J.M.. Mechanisms for initiating cellular DNA replication. Science. 2017; 355:eaah6317. - PubMed
-
- Yao N.Y., O’Donnell M.E. The DNA replication machine: structure and dynamic function. Subcell. Biochem. 2021; 96:233–258. - PubMed
-
- Rajewska M., Wegrzyn K., Konieczny I.. AT-rich region and repeated sequences - the essential elements of replication origins of bacterial replicons. FEMS Microbiol. Rev. 2012; 36:408–434. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

