Microvascular smooth muscle cells exhibit divergent phenotypic switching responses to platelet-derived growth factor and insulin-like growth factor 1
- PMID: 37716411
- PMCID: PMC10842624
- DOI: 10.1016/j.mvr.2023.104609
Microvascular smooth muscle cells exhibit divergent phenotypic switching responses to platelet-derived growth factor and insulin-like growth factor 1
Abstract
Objective: Vascular smooth muscle cell (VSMC) phenotypic switching is critical for normal vessel formation, vascular stability, and healthy brain aging. Phenotypic switching is regulated by mediators including platelet derived growth factor (PDGF)-BB, insulin-like growth factor (IGF-1), as well as transforming growth factor-β (TGF-β) and endothelin-1 (ET-1), but much about the role of these factors in microvascular VSMCs remains unclear.
Methods: We used primary rat microvascular VSMCs to explore PDGF-BB- and IGF-1-induced phenotypic switching.
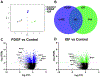
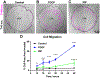
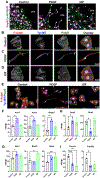
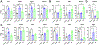
Results: PDGF-BB induced an early proliferative response, followed by formation of polarized leader cells and rapid, directionally coordinated migration. In contrast, IGF-1 induced cell hypertrophy, and only a small degree of migration by unpolarized cells. TGF-β and ET-1 selectively inhibit PDGF-BB-induced VSMC migration primarily by repressing migratory polarization and formation of leader cells. Contractile genes were downregulated by both growth factors, while other genes were differentially regulated by PDGF-BB and IGF-1.
Conclusions: These studies indicate that PDGF-BB and IGF-1 stimulate different types of microvascular VSMC phenotypic switching characterized by different modes of cell migration. Our studies are consistent with a chronic vasoprotective role for IGF-1 in VSMCs in the microvasculature while PDGF is more involved in VSMC proliferation and migration in response to acute activities such as neovascularization. Better understanding of the nuances of the phenotypic switching induced by these growth factors is important for our understanding of a variety of microvascular diseases.
Keywords: Aging; IGF-1; Microvascular VSMC; Microvessels; PDGF; Phenotypic switching; Somatotropic axis; Vascular smooth muscle cell.
Copyright © 2023 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of competing interest Marisa A. Bickel, David M. Sherry, Elizabeth C. Bullen, Michaela L. Vance, Eric W. Howard, and Shannon M. Conley declare no conflict of interest. Ken Jones is CEO for Bioinformatic Solutions, and declares no conflict of interest.
Figures



Similar articles
-
The vasoprotective role of IGF-1 signaling in the cerebral microcirculation: prevention of cerebral microhemorrhages in aging.Geroscience. 2025 Feb;47(1):445-455. doi: 10.1007/s11357-024-01343-5. Epub 2024 Sep 14. Geroscience. 2025. PMID: 39271571 Free PMC article. Review.
-
Chicoric acid prevents PDGF-BB-induced VSMC dedifferentiation, proliferation and migration by suppressing ROS/NFκB/mTOR/P70S6K signaling cascade.Redox Biol. 2018 Apr;14:656-668. doi: 10.1016/j.redox.2017.11.012. Epub 2017 Nov 16. Redox Biol. 2018. PMID: 29175753 Free PMC article.
-
Inhibition of p90RSK Ameliorates PDGF-BB-Mediated Phenotypic Change of Vascular Smooth Muscle Cell and Subsequent Hyperplasia of Neointima.Int J Mol Sci. 2023 Apr 30;24(9):8094. doi: 10.3390/ijms24098094. Int J Mol Sci. 2023. PMID: 37175802 Free PMC article.
-
Platelet-derived growth factor-BB, insulin-like growth factor-I, and phorbol ester activate different signaling pathways for stimulation of vascular smooth muscle cell migration.Exp Cell Res. 1998 Aug 1;242(2):548-60. doi: 10.1006/excr.1998.4138. Exp Cell Res. 1998. PMID: 9683541
-
Platelet-derived growth factor signaling in mesenchymal cells.Front Biosci (Landmark Ed). 2013 Jan 1;18(1):106-19. doi: 10.2741/4090. Front Biosci (Landmark Ed). 2013. PMID: 23276912 Review.
Cited by
-
The vasoprotective role of IGF-1 signaling in the cerebral microcirculation: prevention of cerebral microhemorrhages in aging.Geroscience. 2025 Feb;47(1):445-455. doi: 10.1007/s11357-024-01343-5. Epub 2024 Sep 14. Geroscience. 2025. PMID: 39271571 Free PMC article. Review.
-
Pomegranate as a natural remedy for gastric ulcers prevention: a review of its gastroprotective mechanisms and pharmacological benefits.Naunyn Schmiedebergs Arch Pharmacol. 2025 Jun;398(6):6675-6690. doi: 10.1007/s00210-025-03822-8. Epub 2025 Jan 31. Naunyn Schmiedebergs Arch Pharmacol. 2025. PMID: 39888366 Review.
-
Vascular smooth muscle cell-specific Igf1r deficiency exacerbates the development of hypertension-induced cerebral microhemorrhages and gait defects.Geroscience. 2024 Jun;46(3):3481-3501. doi: 10.1007/s11357-024-01090-7. Epub 2024 Feb 23. Geroscience. 2024. PMID: 38388918 Free PMC article.
-
IGF1R deficiency in vascular smooth muscle cells impairs myogenic autoregulation and cognition in mice.Front Aging Neurosci. 2024 Feb 15;16:1320808. doi: 10.3389/fnagi.2024.1320808. eCollection 2024. Front Aging Neurosci. 2024. PMID: 38425784 Free PMC article.
-
Interplay Between Vascular Dysfunction and Neurodegenerative Pathology: New Insights into Molecular Mechanisms and Management.Biomolecules. 2025 May 13;15(5):712. doi: 10.3390/biom15050712. Biomolecules. 2025. PMID: 40427605 Free PMC article. Review.
References
-
- Savji N., et al., Association between advanced age and vascular disease in different arterial territories: a population database of over 3.6 million subjects. J Am Coll Cardiol, 2013. 61(16): p. 1736–43. - PubMed
-
- Farina FM, et al., miR-128-3p Is a Novel Regulator of Vascular Smooth Muscle Cell Phenotypic Switch and Vascular Diseases. Circ Res, 2020. 126(12): p. e120–e135. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

