Heat shock factor binding protein BrHSBP1 regulates seed and pod development in Brassica rapa
- PMID: 37719218
- PMCID: PMC10499616
- DOI: 10.3389/fpls.2023.1232736
Heat shock factor binding protein BrHSBP1 regulates seed and pod development in Brassica rapa
Abstract
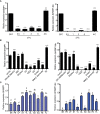
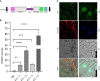
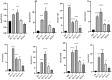
Plant heat shock factor binding proteins (HSBPs) are well known for their implication in the negative regulation of heat stress response (HSR) pathways. Herein, we report on the hitherto unknown functions of HSBP1 in Brassica rapa (BrHSBP1). BrHBSP1 was found to be predominant in flower buds and young leaves, while its segmental duplicate, BrHSBP1-like, was abundant in green siliques. Exposure to abiotic stress conditions, such as heat, drought, cold, and H2O2, and to phytohormones was found to differentially regulate BrHSBP1. The activity of BrHSBP1-GFP fusion proteins revealed their cellular localization in nuclei and cytosols. Transgenic overexpression of BrHSBP1 (BrHSBP1OX) improved pod and seed sizes, while CRISPR-Cas BrHSBP1 knock-out mutants (Brhsbp1_KO) were associated with aborted seed and pod development. The transcriptomic signatures of BrHSBP1OX and Brhsbp1_KO lines revealed that 360 and 2381 genes, respectively, were differentially expressed (Log2FC≥2, padj<0.05) expressed relative to control lines. In particular, developmental processes, including plant reproductive structure development (RSD)-related genes, were relatively downregulated in Brhsbp1_KO. Furthermore, yeast two-hybrid assays confirmed that BrHSBP1 can physically bind to RSD and other genes. Taking the findings together, it is clear that BrHSBP1 is involved in seed development via the modulation of RSD genes. Our findings represent the addition of a new regulatory player in seed and pod development in B. rapa.
Keywords: BrHBSP1; Brassica rapa; CRISPR-Cas; drought; floral genes; heat stress; seed.
Copyright © 2023 Muthusamy, Son, Park and Lee.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
BrEXLB1, a Brassica rapa Expansin-Like B1 Gene is Associated with Root Development, Drought Stress Response, and Seed Germination.Genes (Basel). 2020 Apr 8;11(4):404. doi: 10.3390/genes11040404. Genes (Basel). 2020. PMID: 32276441 Free PMC article.
-
BrPP5.2 Overexpression Confers Heat Shock Tolerance in Transgenic Brassica rapa through Inherent Chaperone Activity, Induced Glucosinolate Biosynthesis, and Differential Regulation of Abiotic Stress Response Genes.Int J Mol Sci. 2021 Jun 16;22(12):6437. doi: 10.3390/ijms22126437. Int J Mol Sci. 2021. PMID: 34208567 Free PMC article.
-
Global Gene-Expression Analysis to Identify Differentially Expressed Genes Critical for the Heat Stress Response in Brassica rapa.PLoS One. 2015 Jun 23;10(6):e0130451. doi: 10.1371/journal.pone.0130451. eCollection 2015. PLoS One. 2015. PMID: 26102990 Free PMC article.
-
Genome-wide analysis of spatiotemporal gene expression patterns during floral organ development in Brassica rapa.Mol Genet Genomics. 2019 Dec;294(6):1403-1420. doi: 10.1007/s00438-019-01585-5. Epub 2019 Jun 20. Mol Genet Genomics. 2019. PMID: 31222475
-
Identification and expression analysis of WRKY family genes under biotic and abiotic stresses in Brassica rapa.Mol Genet Genomics. 2015 Feb;290(1):79-95. doi: 10.1007/s00438-014-0898-1. Epub 2014 Aug 23. Mol Genet Genomics. 2015. PMID: 25149146
Cited by
-
Whole-genome identification of HSF family genes in Cerasus humilis and expression analysis under high-temperature stress.Front Plant Sci. 2025 Apr 28;16:1553187. doi: 10.3389/fpls.2025.1553187. eCollection 2025. Front Plant Sci. 2025. PMID: 40357146 Free PMC article.
References
-
- Augustine S. M., Narayan J. A., Syamaladevi D. P., Appunu C., Chakravarthi M., Ravichandran V., et al. (2015). Erianthus arundinaceus HSP70 (EaHSP70) overexpression increases drought and salinity tolerance in sugarcane (Saccharum spp. hybrid). Plant Sci. 232, 23–34. doi: 10.1016/j.plantsci.2014.12.012 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

