Gene network inference from single-cell omics data and domain knowledge for constructing COVID-19-specific ICAM1-associated pathways
- PMID: 37719701
- PMCID: PMC10501835
- DOI: 10.3389/fgene.2023.1250545
Gene network inference from single-cell omics data and domain knowledge for constructing COVID-19-specific ICAM1-associated pathways
Abstract
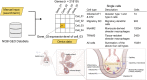
Introduction: Intercellular adhesion molecule 1 (ICAM-1) is a critical molecule responsible for interactions between cells. Previous studies have suggested that ICAM-1 triggers cell-to-cell transmission of HIV-1 or HTLV-1, that SARS-CoV-2 shares several features with these viruses via interactions between cells, and that SARS-CoV-2 cell-to-cell transmission is associated with COVID-19 severity. From these previous arguments, it is assumed that ICAM-1 can be related to SARS-CoV-2 cell-to-cell transmission in COVID-19 patients. Indeed, the time-dependent change of the ICAM-1 expression level has been detected in COVID-19 patients. However, signaling pathways that consist of ICAM-1 and other molecules interacting with ICAM-1 are not identified in COVID-19. For example, the current COVID-19 Disease Map has no entry for those pathways. Therefore, discovering unknown ICAM1-associated pathways will be indispensable for clarifying the mechanism of COVID-19. Materials and methods: This study builds ICAM1-associated pathways by gene network inference from single-cell omics data and multiple knowledge bases. First, single-cell omics data analysis extracts coexpressed genes with significant differences in expression levels with spurious correlations removed. Second, knowledge bases validate the models. Finally, mapping the models onto existing pathways identifies new ICAM1-associated pathways. Results: Comparison of the obtained pathways between different cell types and time points reproduces the known pathways and indicates the following two unknown pathways: (1) upstream pathway that includes proteins in the non-canonical NF-κB pathway and (2) downstream pathway that contains integrins and cytoskeleton or motor proteins for cell transformation. Discussion: In this way, data-driven and knowledge-based approaches are integrated into gene network inference for ICAM1-associated pathway construction. The results can contribute to repairing and completing the COVID-19 Disease Map, thereby improving our understanding of the mechanism of COVID-19.
Keywords: COVID-19; ICAM-1; cell-to-cell transmission; data-driven and knowledge-based approach; model validation; pathway enrichment analysis; single-cell omics data analysis.
Copyright © 2023 Odaka, Magnin and Inoue.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Cardiovascular Risk After SARS-CoV-2 Infection Is Mediated by IL18/IL18R1/HIF-1 Signaling Pathway Axis.Front Immunol. 2022 Jan 5;12:780804. doi: 10.3389/fimmu.2021.780804. eCollection 2021. Front Immunol. 2022. PMID: 35069552 Free PMC article.
-
SARS-CoV-2 attachment to host cells is possibly mediated via RGD-integrin interaction in a calcium-dependent manner and suggests pulmonary EDTA chelation therapy as a novel treatment for COVID 19.Immunobiology. 2021 Jan;226(1):152021. doi: 10.1016/j.imbio.2020.152021. Epub 2020 Nov 5. Immunobiology. 2021. PMID: 33232865 Free PMC article.
-
Network pharmacology and experimental validation identify the potential mechanism of sophocarpine for COVID-19.J Med Microbiol. 2022 May;71(5). doi: 10.1099/jmm.0.001538. J Med Microbiol. 2022. PMID: 35622496
-
Multi-omics approach to COVID-19: a domain-based literature review.J Transl Med. 2021 Dec 7;19(1):501. doi: 10.1186/s12967-021-03168-8. J Transl Med. 2021. PMID: 34876157 Free PMC article. Review.
-
Regulation of intercellular adhesion molecule-1 (CD54) gene expression.J Leukoc Biol. 1999 Dec;66(6):876-88. doi: 10.1002/jlb.66.6.876. J Leukoc Biol. 1999. PMID: 10614768 Review.
References
-
- Benjamini Y., Hochberg Y. (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300. 10.1111/j.2517-6161.1995.tb02031.x - DOI
-
- Blondel V. D., Guillaume J.-L., Lambiotte R., Lefebvre E. (2008). Fast unfolding of communities in large networks. J. Stat. Mech. 2008, P10008. 10.1088/1742-5468/2008/10/p10008 - DOI
LinkOut - more resources
Full Text Sources
Miscellaneous

