Base editing corrects the common Salla disease SLC17A5 c.115C>T variant
- PMID: 37727271
- PMCID: PMC10506058
- DOI: 10.1016/j.omtn.2023.08.024
Base editing corrects the common Salla disease SLC17A5 c.115C>T variant
Abstract
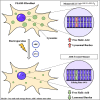
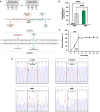
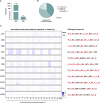
Free sialic acid storage disorders (FSASDs) result from pathogenic variations in the SLC17A5 gene, which encodes the lysosomal transmembrane protein sialin. Loss or deficiency of sialin impairs FSA transport out of the lysosome, leading to cellular dysfunction and neurological impairment, with the most severe form of FSASD resulting in death during early childhood. There are currently no therapies for FSASDs. Here, we evaluated the efficacy of CRISPR-Cas9-mediated homology directed repair (HDR) and adenine base editing (ABE) targeting the founder variant, SLC17A5 c.115C>T (p.Arg39Cys) in human dermal fibroblasts. We observed minimal correction of the pathogenic variant in HDR samples with a high frequency of undesired insertions/deletions (indels) and significant levels of correction for ABE-treated samples with no detectable indels, supporting previous work showing that CRISPR-Cas9-mediated ABE outperforms HDR. Furthermore, ABE treatment of either homozygous or compound heterozygous SLC17A5 c.115C>T human dermal fibroblasts demonstrated significant FSA reduction, supporting amelioration of disease pathology. Translation of this ABE strategy to mouse embryonic fibroblasts harboring the Slc17a5 c.115C>T variant in homozygosity recapitulated these results. Our study demonstrates the feasibility of base editing as a therapeutic approach for the FSASD variant SLC17A5 c.115C>T and highlights the usefulness of base editing in monogenic diseases where transmembrane protein function is impaired.
Keywords: CRISPR-Cas9; MT: RNA/DNA editing; SLC17A5; Salla; free sialic acid storage disease.
© 2023 The Author(s).
Conflict of interest statement
There are no competing interests to disclose.
Figures