Spo13/MEIKIN ensures a Two-Division meiosis by preventing the activation of APC/CAma1 at meiosis I
- PMID: 37728253
- PMCID: PMC10577557
- DOI: 10.15252/embj.2023114288
Spo13/MEIKIN ensures a Two-Division meiosis by preventing the activation of APC/CAma1 at meiosis I
Abstract
Genome haploidization at meiosis depends on two consecutive nuclear divisions, which are controlled by an oscillatory system consisting of Cdk1-cyclin B and the APC/C bound to the Cdc20 activator. How the oscillator generates exactly two divisions has been unclear. We have studied this question in yeast where exit from meiosis involves accumulation of the APC/C activator Ama1 at meiosis II. We show that inactivation of the meiosis I-specific protein Spo13/MEIKIN results in a single-division meiosis due to premature activation of APC/CAma1 . In the wild type, Spo13 bound to the polo-like kinase Cdc5 prevents Ama1 synthesis at meiosis I by stabilizing the translational repressor Rim4. In addition, Cdc5-Spo13 inhibits the activity of Ama1 by converting the B-type cyclin Clb1 from a substrate to an inhibitor of Ama1. Cdc20-dependent degradation of Spo13 at anaphase I unleashes a feedback loop that increases Ama1's synthesis and activity, leading to irreversible exit from meiosis at the second division. Thus, by repressing the exit machinery at meiosis I, Cdc5-Spo13 ensures that cells undergo two divisions to produce haploid gametes.
Keywords: Spo13/MEIKIN; cyclin phosphorylation; meiotic exit; polo-like kinase; translational repression.
© 2023 The Authors. Published under the terms of the CC BY 4.0 license.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
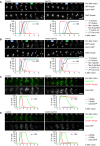
- A, B
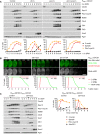
Imaging of spindles (GFP‐tubulin), nucleolar release of Cdc14‐GFP, and TetR‐Tomato, which labels the nucleoplasm (diffuse signal) and the centromeres of one copy of chromosome V (dots). Top, time‐lapse series. First (blue) and second (green) Cdc14 release are marked. Bottom, meiotic events quantified in cells synchronized in silico to spindle formation at metaphase I (t = 0). (A) spo13Δ cells show a longer metaphase I (66 ± 21 min) than control cells (26 ± 9 min, P < 0.0001) and a single round of spindle formation and Cdc14 release. (B) mad2Δ spo13Δ cells show a short metaphase I (10 ± 5 min), similar to mad2Δ cells (10 ± 8 min, P = 0.68), and undergo two rounds of spindle formation and Cdc14 release. Cells in (A) and (B) were filmed together.
- C, D
Imaging of Cdc5‐GFP and SPBs (Cnm67‐Tomato). Top, time‐lapse series. Bottom, the presence of Cdc5 and SPB reduplication quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0). (C) While control cells degrade Cdc5 at meiosis II (four SPBs), spo13Δ cells do so at meiosis I (two SPBs). (D) mad2Δ and mad2Δ spo13Δ cells degrade Cdc5 at meiosis II (four SPBs). Cells in (C) and (D) were filmed together.

Deletion of SPO13 causes Ama1‐dependent proteolysis and nuclear division in P SCC1 ‐CDC20 cells. Top, immunoblot detection of proteins. C, sample from proliferating cells. Bottom, progression of meiosis quantified in fixed cells.
Spo13‐mD reduces Ama1 accumulation and Ama1‐dependent proteolysis at meiosis II. Top, immunoblot detection of proteins. Bottom, relative signal intensities of proteins.
Analysis of mRNA and protein encoded by the ama1 1‐30 ‐mNG locus in P HSL1 ‐CDC20 control and spo13Δ cells. Top, immunoblot detection of proteins. Bottom, mRNA levels relative to the ACT1 transcript were quantified by RT‐qPCR and the 2−ΔCt method. Signal intensities of proteins were normalized to Act1/actin. While control and spo13Δ cells produce ama1 1‐30 ‐mNG mRNA, only spo13Δ cells synthesize mNG protein.
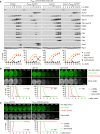
- A
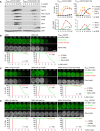
Auxin‐inducible degradation of Rim4 causes Ama1‐dependent proteolysis in P HSL1 ‐CDC20 cells. At t = 7.5 h (arrows), CuSO4 and the auxin 5‐Ph‐IAA were added to induce expression and activation of OsTir1, resulting in degradation of AID‐tagged Rim4‐mNG. Top, immunoblot detection of proteins. Bottom, progression of meiosis quantified in fixed cells.
- B, C
Imaging of Rim4‐mNG and SPBs (Cnm67‐Tomato). Top, time‐lapse series. Bottom, the presence of Rim4‐mNG and SPB reduplication quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0). (B) While control cells degrade Rim4 at meiosis II (four SPBs), spo13Δ mutants do so at meiosis I (two SPBs). Spo13‐mD delays Rim4 degradation by 64 min (95% CI, 58–69 min; P < 0.0001, Welch's t‐test). (C) Deletion of SPO13 causes P HSL1 ‐CDC20 cells to degrade Rim4.
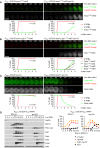
Deletion of SPO13 causes AMA1 translation in Cdc20‐depleted cells. Top, time‐lapse series from the imaging of mNG and SPBs (Cnm67‐Tomato) in P HSL1 ‐CDC20 control and spo13Δ cells carrying ama1 1‐30 ‐mNG in place of AMA1. Bottom, the presence of mNG quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0).
Sporulation genes showing Spo13‐dependent translational repression in metaphase I‐arrested P HSL1 ‐CDC20 ama1Δ strains. Top, time‐lapse series from the imaging of mNG and SPBs (Cnm67‐Tomato) in control and spo13Δ cells carrying gip1 1‐15 ‐mNG in place of GIP1. Bottom, the presence of mNG quantified in cells in which mNG replaces coding sequences downstream of codon 15 of GAT4, GIP1, SPS4, or SSP2. Cells have been synchronized in silico to SPB separation at metaphase I (t = 0).
Deletion of SPO13 causes Rim4 degradation in metaphase I‐arrested P HSL1 ‐CDC20 ama1Δ cells. Top, time‐lapse series from the imaging of Rim4‐mNG and SPBs (Cnm67‐Tomato). Bottom, the presence of Rim4‐mNG quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0).
Deletion of SPO13 causes degradation of APC/C substrates and nuclear division in Cdc20‐depleted (P SCC1 ‐CDC20) cells expressing AMA1 from an estradiol‐inducible GAL promoter (P EST ‐AMA1) at t = 7 h in SPM (arrows). Left, immunoblot detection of proteins. Right, progression of meiosis quantified in fixed cells.

Inhibition of Ime2 activity prevents Rim4 degradation elicited by the SPO13 deletion in metaphase I‐arrested P HSL1 ‐CDC20 ama1Δ cells. Ime2‐as was inhibited with 1Na‐PP1 at 7 h in SPM. Top, time‐lapse series from the imaging of Rim4‐mNG and SPBs (Cnm67‐Tomato). Bottom, the presence of Rim4‐mNG quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0).
Inhibition of Ime2 activity blocks degradation of APC/C substrates and nuclear division in Cdc20‐depleted spo13Δ cells. P EST ‐NDT80 P HSL1 ‐CDC20 strains were released from prophase with estradiol (EST, t = 0), and Ime2‐as was inhibited at metaphase I (1Na‐PP1 at t = 75 min, arrows). Top, immunoblot detection of proteins. Bottom, progression of meiosis quantified in fixed cells.
Ime2‐ΔC induces Rim4 degradation in metaphase I‐arrested P HSL1 ‐CDC20 ama1Δ cells even when Cdc5 is inhibited. Cdc5‐as was inhibited with CMK at 8 h in SPM. Top, time‐lapse series from the imaging of Rim4‐mNG and SPBs (Cnm67‐Tomato). Bottom, the presence of Rim4‐mNG quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0). Rim4 is degraded with similar timing in ime2‐ΔC cells with active or inactive Cdc5 (P = 0.88, Welch's t‐test).
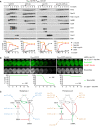
Inhibition of Cdc5 activity blocks degradation of APC/C substrates and nuclear division in Cdc20‐depleted spo13Δ cells. P EST ‐NDT80 P HSL1 ‐CDC20 strains were released from prophase with estradiol (EST, t = 0), and Cdc5‐as was inhibited at metaphase I (CMK at t = 75 min, arrows). Top, immunoblot detection of proteins. Bottom, progression of meiosis quantified in fixed cells.
Cdc5 activity is required for Ime2‐dependent degradation of Rim4. ndt80Δ spo13Δ strains were allowed to arrest at prophase. At 4 h in SPM (t = 0), P EST ‐CDC5 was induced with EST, and Ime2‐as was inhibited with 1Na‐PP1. Top, time‐lapse series from the imaging of Rim4‐mNG and SPBs (Cnm67‐Tomato). Bottom, quantification of the presence of Rim4‐mNG.
The regulatory network controlling Ama1 synthesis and activity. Left, the low‐Ama1/high‐Cdk1 state of metaphase I. Right, the high‐Ama1/low‐Cdk1 state that induces premature exit from metaphase I. Green arrows: activation. Dashed, green arrow: latent activation overwhelmed by inhibition. Red, bar‐headed lines: inhibition. Gray items are inactive or absent. P denotes phosphorylation. Blue, mutations that stabilize metaphase I. Ochre, mutations causing exit from metaphase I. See text for details.

- A
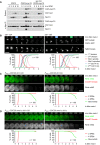
Meiosis in control and spo13‐m2 cells. Top, imaging of spindles (GFP‐tubulin), nucleolar release of Cdc14‐GFP, and TetR‐Tomato, which labels the nucleoplasm (diffuse signal) and the URA3 locus of one copy of chromosome V (dots). First (blue) and second (green) Cdc14 release are marked. Bottom, meiotic events quantified in cells synchronized in silico to spindle formation at metaphase I (t = 0). spo13‐m2 prolongs metaphase I to 60 ± 26 min (control, 23 ± 6 min; P < 0.0001, Welch's t‐test) and causes ~38% of cells to undergo a single round of spindle formation and Cdc14 release.
- B
spo13‐m2 causes Ama1‐dependent proteolysis and nuclear division in P HSL1 ‐CDC20 cells. Top, immunoblot detection of proteins. Bottom, progression of meiosis quantified in fixed cells.
- C
Imaging of Rim4‐mNG and SPBs (Cnm67‐Tomato) in P HSL1 ‐CDC20 cells carrying spo13‐m2 or cdc5‐as. Cdc5‐as was inhibited with CMK at 7.5 h in SPM. Top, time‐lapse series. Bottom, the presence of Rim4‐mNG quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0).
- D, E
Extracts from metaphase I‐arrested P HSL1 ‐CDC20 cells (8 h in SPM) were applied to columns carrying no antibody or antibodies to either Spo13 or Cdc5. Flow‐throughs were analyzed by immunoblotting. (D) Depletion of Spo13 from extracts of control cells (SPO13) removes little Cdc5 (10%). (E) Depletion of Spo13 from extracts of cells overexpressing Spo13 (P EST ‐SPO13, estradiol for 45 min) removes most of Cdc5 (97%).
Deletion of CLB1 causes Ama1‐dependent proteolysis and nuclear division in P HSL1 ‐CDC20 cells. Top, immunoblot detection of proteins. Bottom, progression of meiosis quantified in fixed cells.
Imaging of Rim4‐mNG and SPBs (Cnm67‐Tomato) in P HSL1 ‐CDC20 cells carrying clb1Δ, cdk1‐as2, or spo13‐10A. Cdk1‐as2 was inhibited with 1Na‐PP1 at 7.5 h in SPM. Top, time‐lapse series. Bottom, the presence of Rim4‐mNG quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0).
Inhibition of Cdk1 at metaphase I causes accumulation of Ama1 and degradation of APC/C substrates. P EST ‐NDT80 P HSL1 ‐CDC20 strains were released from prophase with estradiol (EST, t = 0), and Cdk1‐as2 was inhibited at metaphase I (1Na‐PP1 at t = 120 min, arrows). Left, immunoblot detection of proteins. Right, progression of meiosis quantified in fixed cells.

- A–C
Imaging of Rim4‐mNG and SPBs (Cnm67‐Tomato) in metaphase I‐arrested P HSL1 ‐CDC20 ama1Δ strains. Top, time‐lapse series. Bottom, the presence of mNG quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0). (A) Inhibition of Hrr25 activity prevents Rim4 degradation elicited by the SPO13 deletion. Hrr25‐as was inhibited with 1NM‐PP1 at 7 h in SPM. (B) Inhibition of Hrr25 prevents Rim4 degradation induced by hyperactive Ime2‐ΔC. Hrr25‐as was inhibited with 1NM‐PP1 at 7 h in SPM. (C) Rim4 is degraded with similar timing in hrr25‐ΔC cells with active or inactive Ime2 (P = 0.91, Welch's t‐test). Ime2‐as was inhibited with 1Na‐PP1 at 7 h in SPM.
- D
Binding of Rim4‐mNG and Ime2 to Hrr25‐myc9 immunoprecipitated with α‐Myc antibodies from extracts of P HSL1 ‐CDC20 cells.
- A
Ama1‐6A causes degradation of APC/C substrates and nuclear division in P HSL1 ‐CDC20 cells. Left, immunoblot detection of proteins. Top right, progression of meiosis quantified in fixed cells. Bottom right, relative signal intensities of proteins.
- B–D
Analysis of Rim4 degradation in P HSL1 ‐CDC20 cells carrying AMA1‐6A. Top, time‐lapse series from the filming of Rim4‐mNG and SPBs (Cnm67‐Tomato). Bottom, the presence of Rim4‐mNG quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0). (B) Ama1‐6A causes Rim4 degradation. (C) Rim4 degradation is not affected by either spo13‐mD (P = 0.86) or P DMC1 ‐clb1‐mDK (P = 0.20) alone, but is blocked in the presence of both alleles (P < 0.0001, Welch's t‐test). Cells in (B) and (C) were filmed together. (D) Rim4 degradation depends on the activities of Cdc5, Ime2, and Hrr25. Analogue‐sensitive kinases were inhibited at 8 h in SPM.

- A–C
Binding of cyclins to the APC/C immunoprecipitated with α‐Apc2 antibodies. (A) Clb1 and Clb3 but not the other Clbs copurify with the APC/C. (B) The degrons of Clb1 are not required for binding to the APC/C. Ha3‐tagged Clb1 and Clb1‐mDK were expressed from the DMC1 promoter. (C) Deletion of AMA1 reduces Clb1's interaction with the APC/C.
- D
Proteins interacting with Ama1 at metaphase I. Ama1‐GFP and Esp1‐GFP (negative control) were purified from P HSL1 ‐CDC20 strains at 8 h in SPM, digested with trypsin and Lys‐C, and subjected to LC–MS/MS. ‐log10‐transformed P‐values and mean log2‐transformed label‐free quantifications of copurifying proteins were obtained from MaxQuant and displayed as a volcano plot. Baits (green), selected proteins with P < 0.05 (red) or P > 0.05 (orange), and the eight subunits of the CCT chaperonin (brown) are labeled. CCT is known to encapsulate and fold APC/C activators (Camasses et al, 2003). Only a subset of APC/C subunits is marked. Note that C‐terminally tagged Ama1 cannot bind the APC/C.
- E
Binding of Clb1, Clb3, and Cdk1 to Ama1‐myc9 immunoprecipitated with α‐Myc antibodies.
Clb1‐ha3 binds to Cdc15‐myc15 in α‐Myc immunoprecipitations from extracts of control and spo13Δ cells.
Meiosis in clb1‐12A and clb1‐12D cells. Top, time lapse series from the imaging of spindles (GFP‐tubulin), nucleolar release of Cdc14‐GFP, and TetR‐Tomato, which labels the nucleoplasm (diffuse signal) and the centromeres of one copy of chromosome V (dots). First (blue) and second (green) Cdc14 release are marked. Bottom, meiotic events quantified in cells synchronized in silico to spindle formation at metaphase I (t = 0). The duration of metaphase I is similar in clb1‐12A and clb1‐12D cells (P = 0.33).
P HSL1 ‐CDC20 clb1‐12A cells degrade Rim4 earlier than P HSL1 ‐CDC20 clb1‐6A cells (P < 0.0001). Top, time‐lapse series from the imaging of Rim4‐mNG and SPBs (Cnm67‐Tomato). Bottom, the presence of Rim4‐mNG quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0).
Clb1 phosphorylation is not required for the activation of Spo13. Top, time‐lapse series from the imaging of Rim4‐mNG and SPBs (Cnm67‐Tomato) in P HSL1 ‐CDC20 ama1Δ cells carrying clb1‐12A or clb1Δ. Bottom, the presence of Rim4‐mNG quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0).
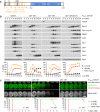
Domain structure of Clb1 showing D‐ and KEN‐boxes (black), phospho‐serines (orange), and cyclin boxes (blue).
clb1‐12A but not clb1‐12D causes Ama1‐dependent proteolysis and nuclear division in Cdc20‐depleted cells. Top, immunoblot detection of proteins. Bottom, progression of meiosis quantified in fixed cells.
Imaging of Rim4‐mNG and SPBs (Cnm67‐Tomato) in P HSL1 ‐CDC20 control cells and cells carrying clb1‐12A or clb1‐12D. Top, time‐lapse series. Bottom, the presence of Rim4‐mNG quantified in cells synchronized in silico to SPB separation at metaphase I (t = 0).

Stability of Clb1, Clb1‐12A, and Clb1‐12D. CLB1 alleles were expressed from the DMC1 promoter in ndt80Δ cdk1‐as1 strains carrying AMA1 or ama1Δ. Strains were arrested at prophase in the presence of 1NM‐PP1 to inhibit Cdk1‐as1. At 6 h in SPM, translation was inhibited with cycloheximide (CHX, t = 0). Left, immunoblot detection of Clb1. Right, log2‐transformed Clb1 signal intensities (normalized to t = 0) plotted over time in CHX. Half‐lives were calculated by linear regression.
Analysis of APC/C substrates in P SCC1 ‐CDC20 cells carrying spo13Δ and CLB1, clb1‐12A, or clb1‐12D. Top, immunoblot detection of proteins. Bottom, progression of meiosis quantified in fixed cells.
Spo13 stability at prophase. ndt80Δ cdk1‐as1 strains carrying AMA1 or ama1Δ were arrested at prophase in the presence of 1NM‐PP1 to inhibit Cdk1‐as1. At 6 h in SPM, translation was inhibited with CHX (t = 0). mnd2Δ cells expressing Spo13 from the DMC1 promoter were processed in parallel. Left, immunoblot detection of Spo13. Right, log2‐transformed Spo13 signal intensities (normalized to t = 0) plotted over time in CHX. Half‐lives were calculated by linear regression. Open diamond, data point omitted from regression.
References
-
- Arguello‐Miranda O, Zagoriy I, Mengoli V, Rojas J, Jonak K, Oz T, Graf P, Zachariae W (2017) Casein kinase 1 coordinates cohesin cleavage, gametogenesis, and exit from M phase in meiosis II. Dev Cell 40: 37–52 - PubMed
-
- Asakawa H, Kitamura K, Shimoda C (2001) A novel Cdc20‐related WD‐repeat protein, Fzr1, is required for spore formation in Schizosaccharomyces pombe . Mol Genet Genomics 265: 424–435 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

