Formate Supplementation Enhances Antitumor CD8+ T-cell Fitness and Efficacy of PD-1 Blockade
- PMID: 37728660
- PMCID: PMC10843486
- DOI: 10.1158/2159-8290.CD-22-1301
Formate Supplementation Enhances Antitumor CD8+ T-cell Fitness and Efficacy of PD-1 Blockade
Abstract
The tumor microenvironment (TME) restricts antitumor CD8+ T-cell function and immunotherapy responses. Cancer cells compromise the metabolic fitness of CD8+ T cells within the TME, but the mechanisms are largely unknown. Here we demonstrate that one-carbon (1C) metabolism is enhanced in T cells in an antigen-specific manner. Therapeutic supplementation of 1C metabolism using formate enhances CD8+ T-cell fitness and antitumor efficacy of PD-1 blockade in B16-OVA tumors. Formate supplementation drives transcriptional alterations in CD8+ T-cell metabolism and increases gene signatures for cellular proliferation and activation. Combined formate and anti-PD-1 therapy increases tumor-infiltrating CD8+ T cells, which are essential for enhanced tumor control. Our data demonstrate that formate provides metabolic support to CD8+ T cells reinvigorated by anti-PD-1 to overcome a metabolic vulnerability in 1C metabolism in the TME to further improve T-cell function.
Significance: This study identifies that deficiencies in 1C metabolism limit the efficacy of PD-1 blockade in B16-OVA tumors. Supplementing 1C metabolism with formate during anti-PD-1 therapy enhances CD8+ T-cell fitness in the TME and CD8+ T-cell-mediated tumor clearance. These findings demonstrate that formate supplementation can enhance exhausted CD8+ T-cell function. See related commentary by Lin et al., p. 2507. This article is featured in Selected Articles from This Issue, p. 2489.
©2023 American Association for Cancer Research.
Conflict of interest statement
Competing interests
AHS has patents/pending royalties on the PD-1 pathway from Roche and Novartis. AHS is on advisory boards for SQZ Biotechnologies, Elpiscience, Selecta, Bicara, Monopteros, Fibrogen, IOME, Alixia, GlaxoSmithKline, Janssen and Amgen. She also is on scientific advisory boards for the Massachusetts General Cancer Center, Program in Cellular and Molecular Medicine at Boston Children’s Hospital, the Human Oncology and Pathogenesis Program at Memorial Sloan Kettering Cancer Center, the Gladstone Institute, and the Johns Hopkins Bloomberg-Kimmel Institute for Cancer Immunotherapy. AHS has received research funding from Merck, Vertex, Moderna, Quark/Iome, Erasca and AbbVie unrelated to this project. GJF has patents/pending royalties on the PD-L1/PD-1 pathway from Roche, Merck MSD, Bristol-Myers-Squibb, Merck KGA, Boehringer-Ingelheim, AstraZeneca, Dako, Leica, Mayo Clinic, Eli Lilly, and Novartis. GJF has served on advisory boards for Roche, Bristol-Myers-Squibb, Origimed, Triursus, iTeos, NextPoint, IgM, Jubilant, Trillium, GV20, IOME, and Geode. GJF has equity in Nextpoint, Triursus, Xios, iTeos, IgM, Trillium, Invaria, GV20, and Geode. MCH is on advisory boards for MitoQ, Alixia, and Minovia. MCH has received research funding from Roche and Agilent.
Figures





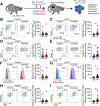
Comment in
-
Re-"Formate" T-cell Antitumor Responses.Cancer Discov. 2023 Dec 12;13(12):2507-2509. doi: 10.1158/2159-8290.CD-23-1059. Cancer Discov. 2023. PMID: 38084093
References
-
- Chen DS, Mellman I. Oncology meets immunology: the cancer-immunity cycle. Immunity; 2013;39:1–10. Available from: https://pubmed.ncbi.nlm.nih.gov/23890059/ - PubMed
-
- Baumeister SH, Freeman GJ, Dranoff G, Sharpe AH. Coinhibitory Pathways in Immunotherapy for Cancer. Annu Rev Immunol; 2016;34:539–73. Available from: https://pubmed.ncbi.nlm.nih.gov/26927206/ - PubMed
-
- Wolchok JD, Kluger H, Callahan MK, Postow MA, Rizvi NA, Lesokhin AM, et al. Nivolumab plus ipilimumab in advanced melanoma. N Engl J Med; 2013;369:122–33. Available from: https://pubmed.ncbi.nlm.nih.gov/23724867/ - PMC - PubMed
-
- Postow MA, Chesney J, Pavlick AC, Robert C, Grossmann K, McDermott D, et al. Nivolumab and ipilimumab versus ipilimumab in untreated melanoma. N Engl J Med; 2015;372:2006–17. Available from: https://pubmed.ncbi.nlm.nih.gov/25891304/ - PMC - PubMed
-
- Carbone DP, Reck M, Paz-Ares L, Creelan B, Horn L, Steins M, et al. First-Line Nivolumab in Stage IV or Recurrent Non-Small-Cell Lung Cancer. N Engl J Med; 2017;376:2415–26. Available from: https://pubmed.ncbi.nlm.nih.gov/28636851/ - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

